Acta Agriculturae Zhejiangensis ›› 2026, Vol. 38 ›› Issue (1): 35-53.DOI: 10.3969/j.issn.1004-1524.20250081
• Crop Science • Previous Articles Next Articles
Identification of GRAS gene family members and their regulatory roles in reproductive development in Chenopodium quinoa
YANG Yanga( ), ZHANG Shuaia, DONG Chenwenhuaa,b, ZENG Mengqionga, LIN Chuna,b, MAO Zichaoa,b,*(
), ZHANG Shuaia, DONG Chenwenhuaa,b, ZENG Mengqionga, LIN Chuna,b, MAO Zichaoa,b,*( )
)
- College of Agriculture and Biotechnology; b. Research Center of Featured Minor Crops, Yunnan Agricultural University, Kunming 650201, China
-
Received:2025-02-07Online:2026-01-25Published:2026-02-11
CLC Number:
Cite this article
YANG Yang, ZHANG Shuai, DONG Chenwenhua, ZENG Mengqiong, LIN Chun, MAO Zichao. Identification of GRAS gene family members and their regulatory roles in reproductive development in Chenopodium quinoa[J]. Acta Agriculturae Zhejiangensis, 2026, 38(1): 35-53.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.zjnyxb.cn/EN/10.3969/j.issn.1004-1524.20250081
| 引物名称primer name | 引物序列(5'→3') Primer sequence(5'→3') | 产物大小/bp Product length/bp |
|---|---|---|
| 1-CqDELLA01-F2 | CGAAAGACACGAGCCACTAATAAA | 169 |
| 1-CqDELLA01-R2 | CCACCCTAGAGTCAAGCACCC | |
| 2-CqDELLA02-F2 | ACTTCTAGTTCTCAATCTTTTGTTTTC | 194 |
| 2-CqDELLA02-R2 | AAAGTGTCTAGCACCCCCAG | |
| 3-CqPAT1-05-F2 | CTTTCCCTTAGCAATGACAACA | 105 |
| 3-CqPAT1-05-R2 | AATGTTTAGGAAGGATGGTCGT | |
| 4-CqPAT1-06-F | AGGACTACTTCGGTGTTACTCTG | 103 |
| 4-CqPAT1-06-R | CCAAGCTGAAGCCGAGATCAA | |
| 5-CqDLT01-F2 | GGTATCAGGAGGGACGAAAG | 166 |
| 5-CqDLT01-R2 | CAGCAGCATCAACAGAACCC | |
| 6-CqDLT02-F2 | CATGGATTAGTCTTTGATTTTCGAT | 175 |
| 6-CqDLT02-R2 | CATGCCATAAGTAGATCAAATAACAA | |
| 7-CqSHR05-F | GGCGTATAGTGAGGAGGTTTGTG | 109 |
| 7-CqSHR05-R | TATTCCGGCAGTCGGAGAC | |
| 8-CqSHR06-F | CGGAGAGGAGAGAGAATGCC | 137 |
| 8-CqSHR06-R | TGCCATCGACCACCCTTCT | |
| CqActin-F2 | TTGGAATCTCGCCGCCAAA | 132 |
| CqActin-R2 | CCAGGTCCACCAACAATCCA |
Table 1 Sequences of qRT-PCR primers
| 引物名称primer name | 引物序列(5'→3') Primer sequence(5'→3') | 产物大小/bp Product length/bp |
|---|---|---|
| 1-CqDELLA01-F2 | CGAAAGACACGAGCCACTAATAAA | 169 |
| 1-CqDELLA01-R2 | CCACCCTAGAGTCAAGCACCC | |
| 2-CqDELLA02-F2 | ACTTCTAGTTCTCAATCTTTTGTTTTC | 194 |
| 2-CqDELLA02-R2 | AAAGTGTCTAGCACCCCCAG | |
| 3-CqPAT1-05-F2 | CTTTCCCTTAGCAATGACAACA | 105 |
| 3-CqPAT1-05-R2 | AATGTTTAGGAAGGATGGTCGT | |
| 4-CqPAT1-06-F | AGGACTACTTCGGTGTTACTCTG | 103 |
| 4-CqPAT1-06-R | CCAAGCTGAAGCCGAGATCAA | |
| 5-CqDLT01-F2 | GGTATCAGGAGGGACGAAAG | 166 |
| 5-CqDLT01-R2 | CAGCAGCATCAACAGAACCC | |
| 6-CqDLT02-F2 | CATGGATTAGTCTTTGATTTTCGAT | 175 |
| 6-CqDLT02-R2 | CATGCCATAAGTAGATCAAATAACAA | |
| 7-CqSHR05-F | GGCGTATAGTGAGGAGGTTTGTG | 109 |
| 7-CqSHR05-R | TATTCCGGCAGTCGGAGAC | |
| 8-CqSHR06-F | CGGAGAGGAGAGAGAATGCC | 137 |
| 8-CqSHR06-R | TGCCATCGACCACCCTTCT | |
| CqActin-F2 | TTGGAATCTCGCCGCCAAA | 132 |
| CqActin-R2 | CCAGGTCCACCAACAATCCA |
| 基因名称 Gene name | 基因ID Gene ID | 染色体 Chromosome | 氨基酸长度/aa Amino acid length/aa | 蛋白质分子质量/u Protein molecular weight/u | 等电点 Isoelectric point | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|
| CqDELLA01 | CQ012679 | Cq8B | 459 | 50 811.86 | 5.51 | 溶酶体Cytolysosome |
| CqDELLA02 | CQ040570 | Cq8A | 474 | 52 509.87 | 5.66 | 溶酶体Cytolysosome |
| CqDELLA03 | CQ001679 | Cq6B | 617 | 67 367.76 | 4.90 | 细胞核Nuclear |
| CqDELLA04 | CQ026984 | Cq6A | 618 | 67 500.87 | 4.90 | 细胞核Nuclear |
| CqDLT01 | CQ004440 | Cq5B | 648 | 73 031.86 | 5.77 | 叶绿体Chloroplast |
| CqDLT02 | CQ029757 | Cq5A | 657 | 73 968.04 | 5.47 | 叶绿体Chloroplast |
| CqHAM01 | CQ002321 | Cq6B | 610 | 67 095.88 | 6.46 | 细胞核Nuclear |
| CqHAM02 | CQ027701 | Cq6A | 602 | 66 232.26 | 6.82 | 细胞核Nuclear |
| CqHAM03 | CQ003474 | Cq5B | 785 | 85 801.61 | 6.15 | 细胞核Nuclear |
| CqHAM04 | CQ028875 | Cq5A | 781 | 85 891.76 | 6.12 | 细胞核Nuclear |
| CqLAS01 | CQ033480 | Cq9B | 490 | 55 803.58 | 7.02 | 细胞核Nuclear |
| CqLAS02 | CQ051297 | Cq9A | 484 | 54 848.37 | 6.76 | 细胞核Nuclear |
| CqLISCL01 | CQ032605 | Cq9B | 350 | 41 049.14 | 8.10 | 溶酶体Cytolysosome |
| CqLISCL02 | CQ041202 | Cq7A | 725 | 82 353.95 | 4.95 | 细胞核Nuclear |
| CqLISCL03 | CQ032598 | Cq9B | 689 | 78 590.85 | 5.18 | 细胞核Nuclear |
| CqLISCL04 | CQ041204 | Cq7A | 619 | 70 272.91 | 6.29 | 细胞核Nuclear |
| CqLISCL05 | CQ032601 | Cq9B | 679 | 76 337.79 | 5.60 | 细胞核Nuclear |
| CqLISCL06 | CQ032604 | Cq9B | 548 | 62 438.96 | 7.94 | 溶酶体Cytolysosome |
| CqLISCL07 | CQ041201 | Cq7A | 662 | 74 879.34 | 6.61 | 细胞核Nuclear |
| CqLISCL08 | CQ032603 | Cq9B | 667 | 75 299.73 | 5.23 | 细胞核Nuclear |
| CqLISCL09 | CQ041200 | Cq7A | 686 | 77 311.71 | 4.87 | 细胞核Nuclear |
| CqLISCL10 | CQ030634 | Cq5A | 760 | 86 162.35 | 5.50 | 细胞核Nuclear |
| CqLISCL11 | CQ055311 | Cq2A | 780 | 87 684.22 | 7.07 | 溶酶体Cytolysosome |
| CqLISCL12 | CQ025002 | Cq1B | 818 | 91 640.62 | 7.49 | 细胞核Nuclear |
| CqLISCL13 | CQ032609 | Cq9B | 756 | 84 422.57 | 5.55 | 细胞核Nuclear |
| CqLISCL14 | CQ041197 | Cq7A | 756 | 84 533.61 | 5.81 | 细胞核Nuclear |
| CqPAT1-01 | CQ024284 | Cq1B | 545 | 61 454.56 | 4.73 | 叶绿体Chloroplast |
| CqPAT1-02 | CQ048458 | Cq1A | 426 | 48 153.43 | 5.27 | 溶酶体Cytolysosome |
| CqPAT1-03 | CQ016853 | Cq3B | 639 | 70 122.51 | 5.46 | 细胞核Nuclear |
| CqPAT1-04 | CQ044170 | Cq3A | 767 | 84 245.91 | 6.39 | 细胞核Nuclear |
| CqPAT1-05 | CQ022180 | Cq4B | 635 | 70 854.40 | 8.16 | 叶绿体Chloroplast |
| CqPAT1-06 | CQ037875 | Cq4A | 634 | 70 854.39 | 7.57 | 叶绿体Chloroplast |
| CqPAT1-07 | CQ010139 | Cq8B | 544 | 61 079.89 | 6.33 | 细胞核Nuclear |
| CqPAT1-08 | CQ038026 | Cq8A | 544 | 61 110.88 | 6.51 | 细胞核Nuclear |
| CqPAT1-09 | CQ013243 | Cq2B | 466 | 52 280.60 | 5.82 | 叶绿体Chloroplast |
| CqPAT1-10 | CQ034654 | Cq9B | 541 | 60 293.76 | 6.56 | 细胞核Nuclear |
| CqPAT1-11 | CQ050106 | Cq9A | 588 | 65 884.16 | 6.69 | 叶绿体Chloroplast |
| CqSCL3L01 | CQ019876 | Cq4B | 454 | 50 710.01 | 6.51 | 细胞核Nuclear |
| CqSCL3L02 | CQ035148 | Cq4A | 454 | 50 837.30 | 6.65 | 细胞核Nuclear |
| CqSCL4L01 | CQ003361 | Cq6B | 557 | 60 789.24 | 4.38 | 细胞核Nuclear |
| CqSCL4L02 | CQ028754 | Cq6A | 610 | 67 157.51 | 4.74 | 细胞核Nuclear |
| CqSCR01 | CQ052618 | Cq9A | 872 | 95 159.47 | 6.39 | 细胞核Nuclear |
| CqSCR02 | CQ007129 | Cq7B | 861 | 93 906.00 | 6.40 | 细胞核Nuclear |
| CqSCR03 | CQ014997 | Cq2B | 396 | 43 813.31 | 6.99 | 叶绿体Chloroplast |
| CqSCR04 | CQ054542 | Cq2A | 396 | 43 859.22 | 7.32 | 细胞核Nuclear |
| CqSHR01 | CQ003291 | Cq6B | 475 | 53 457.35 | 5.88 | 线粒体、细胞骨架 Mitochondrion, cytoskeleton |
| CqSHR02 | CQ028690 | Cq6A | 389 | 43 994.11 | 5.05 | 细胞骨架Cytoskeleton |
| CqSHR03 | CQ018831 | Cq3B | 439 | 50 575.84 | 6.38 | 叶绿体Chloroplast |
| CqSHR04 | CQ044974 | Cq3A | 440 | 50 599.98 | 6.73 | 叶绿体Chloroplast |
| CqSHR05 | CQ003315 | Cq6B | 472 | 53 064.48 | 5.94 | 细胞核Nuclear |
| CqSHR06 | CQ028714 | Cq6A | 467 | 52 415.84 | 5.86 | 细胞核Nuclear |
Table 2 Information of GRAS gene family in quinoa
| 基因名称 Gene name | 基因ID Gene ID | 染色体 Chromosome | 氨基酸长度/aa Amino acid length/aa | 蛋白质分子质量/u Protein molecular weight/u | 等电点 Isoelectric point | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|
| CqDELLA01 | CQ012679 | Cq8B | 459 | 50 811.86 | 5.51 | 溶酶体Cytolysosome |
| CqDELLA02 | CQ040570 | Cq8A | 474 | 52 509.87 | 5.66 | 溶酶体Cytolysosome |
| CqDELLA03 | CQ001679 | Cq6B | 617 | 67 367.76 | 4.90 | 细胞核Nuclear |
| CqDELLA04 | CQ026984 | Cq6A | 618 | 67 500.87 | 4.90 | 细胞核Nuclear |
| CqDLT01 | CQ004440 | Cq5B | 648 | 73 031.86 | 5.77 | 叶绿体Chloroplast |
| CqDLT02 | CQ029757 | Cq5A | 657 | 73 968.04 | 5.47 | 叶绿体Chloroplast |
| CqHAM01 | CQ002321 | Cq6B | 610 | 67 095.88 | 6.46 | 细胞核Nuclear |
| CqHAM02 | CQ027701 | Cq6A | 602 | 66 232.26 | 6.82 | 细胞核Nuclear |
| CqHAM03 | CQ003474 | Cq5B | 785 | 85 801.61 | 6.15 | 细胞核Nuclear |
| CqHAM04 | CQ028875 | Cq5A | 781 | 85 891.76 | 6.12 | 细胞核Nuclear |
| CqLAS01 | CQ033480 | Cq9B | 490 | 55 803.58 | 7.02 | 细胞核Nuclear |
| CqLAS02 | CQ051297 | Cq9A | 484 | 54 848.37 | 6.76 | 细胞核Nuclear |
| CqLISCL01 | CQ032605 | Cq9B | 350 | 41 049.14 | 8.10 | 溶酶体Cytolysosome |
| CqLISCL02 | CQ041202 | Cq7A | 725 | 82 353.95 | 4.95 | 细胞核Nuclear |
| CqLISCL03 | CQ032598 | Cq9B | 689 | 78 590.85 | 5.18 | 细胞核Nuclear |
| CqLISCL04 | CQ041204 | Cq7A | 619 | 70 272.91 | 6.29 | 细胞核Nuclear |
| CqLISCL05 | CQ032601 | Cq9B | 679 | 76 337.79 | 5.60 | 细胞核Nuclear |
| CqLISCL06 | CQ032604 | Cq9B | 548 | 62 438.96 | 7.94 | 溶酶体Cytolysosome |
| CqLISCL07 | CQ041201 | Cq7A | 662 | 74 879.34 | 6.61 | 细胞核Nuclear |
| CqLISCL08 | CQ032603 | Cq9B | 667 | 75 299.73 | 5.23 | 细胞核Nuclear |
| CqLISCL09 | CQ041200 | Cq7A | 686 | 77 311.71 | 4.87 | 细胞核Nuclear |
| CqLISCL10 | CQ030634 | Cq5A | 760 | 86 162.35 | 5.50 | 细胞核Nuclear |
| CqLISCL11 | CQ055311 | Cq2A | 780 | 87 684.22 | 7.07 | 溶酶体Cytolysosome |
| CqLISCL12 | CQ025002 | Cq1B | 818 | 91 640.62 | 7.49 | 细胞核Nuclear |
| CqLISCL13 | CQ032609 | Cq9B | 756 | 84 422.57 | 5.55 | 细胞核Nuclear |
| CqLISCL14 | CQ041197 | Cq7A | 756 | 84 533.61 | 5.81 | 细胞核Nuclear |
| CqPAT1-01 | CQ024284 | Cq1B | 545 | 61 454.56 | 4.73 | 叶绿体Chloroplast |
| CqPAT1-02 | CQ048458 | Cq1A | 426 | 48 153.43 | 5.27 | 溶酶体Cytolysosome |
| CqPAT1-03 | CQ016853 | Cq3B | 639 | 70 122.51 | 5.46 | 细胞核Nuclear |
| CqPAT1-04 | CQ044170 | Cq3A | 767 | 84 245.91 | 6.39 | 细胞核Nuclear |
| CqPAT1-05 | CQ022180 | Cq4B | 635 | 70 854.40 | 8.16 | 叶绿体Chloroplast |
| CqPAT1-06 | CQ037875 | Cq4A | 634 | 70 854.39 | 7.57 | 叶绿体Chloroplast |
| CqPAT1-07 | CQ010139 | Cq8B | 544 | 61 079.89 | 6.33 | 细胞核Nuclear |
| CqPAT1-08 | CQ038026 | Cq8A | 544 | 61 110.88 | 6.51 | 细胞核Nuclear |
| CqPAT1-09 | CQ013243 | Cq2B | 466 | 52 280.60 | 5.82 | 叶绿体Chloroplast |
| CqPAT1-10 | CQ034654 | Cq9B | 541 | 60 293.76 | 6.56 | 细胞核Nuclear |
| CqPAT1-11 | CQ050106 | Cq9A | 588 | 65 884.16 | 6.69 | 叶绿体Chloroplast |
| CqSCL3L01 | CQ019876 | Cq4B | 454 | 50 710.01 | 6.51 | 细胞核Nuclear |
| CqSCL3L02 | CQ035148 | Cq4A | 454 | 50 837.30 | 6.65 | 细胞核Nuclear |
| CqSCL4L01 | CQ003361 | Cq6B | 557 | 60 789.24 | 4.38 | 细胞核Nuclear |
| CqSCL4L02 | CQ028754 | Cq6A | 610 | 67 157.51 | 4.74 | 细胞核Nuclear |
| CqSCR01 | CQ052618 | Cq9A | 872 | 95 159.47 | 6.39 | 细胞核Nuclear |
| CqSCR02 | CQ007129 | Cq7B | 861 | 93 906.00 | 6.40 | 细胞核Nuclear |
| CqSCR03 | CQ014997 | Cq2B | 396 | 43 813.31 | 6.99 | 叶绿体Chloroplast |
| CqSCR04 | CQ054542 | Cq2A | 396 | 43 859.22 | 7.32 | 细胞核Nuclear |
| CqSHR01 | CQ003291 | Cq6B | 475 | 53 457.35 | 5.88 | 线粒体、细胞骨架 Mitochondrion, cytoskeleton |
| CqSHR02 | CQ028690 | Cq6A | 389 | 43 994.11 | 5.05 | 细胞骨架Cytoskeleton |
| CqSHR03 | CQ018831 | Cq3B | 439 | 50 575.84 | 6.38 | 叶绿体Chloroplast |
| CqSHR04 | CQ044974 | Cq3A | 440 | 50 599.98 | 6.73 | 叶绿体Chloroplast |
| CqSHR05 | CQ003315 | Cq6B | 472 | 53 064.48 | 5.94 | 细胞核Nuclear |
| CqSHR06 | CQ028714 | Cq6A | 467 | 52 415.84 | 5.86 | 细胞核Nuclear |
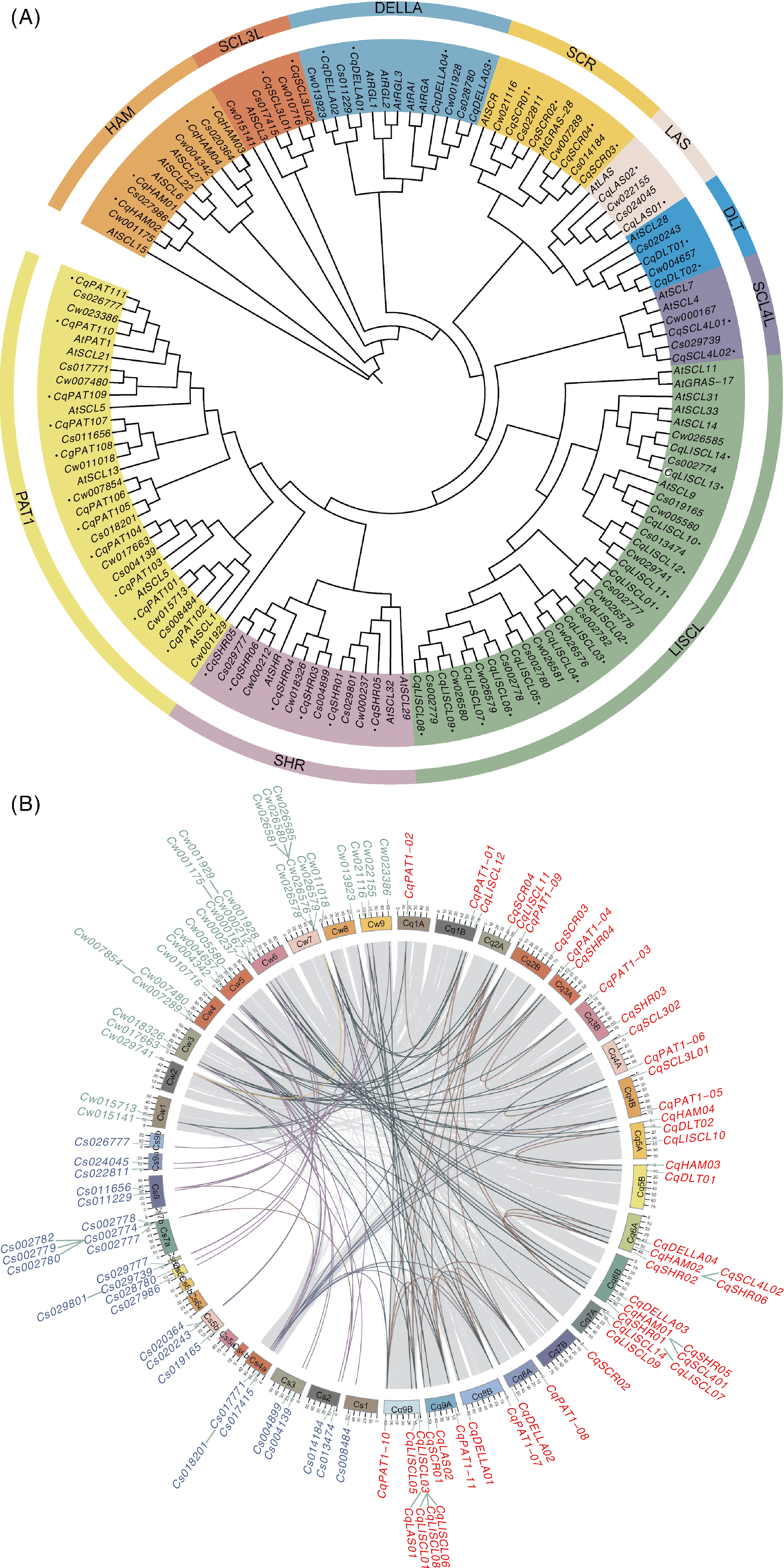
Fig.1 Phylogenetic tree and collinearity analysis of GRAS gene family A, Phylogenetic tree of the GRAS family among C. watsonii, C. suecicum, C. quinoa and Arabidopsis thaliana; B, GRAS gene collinearity analysis among C. watsonii, C. suecicum and C. quinoa.
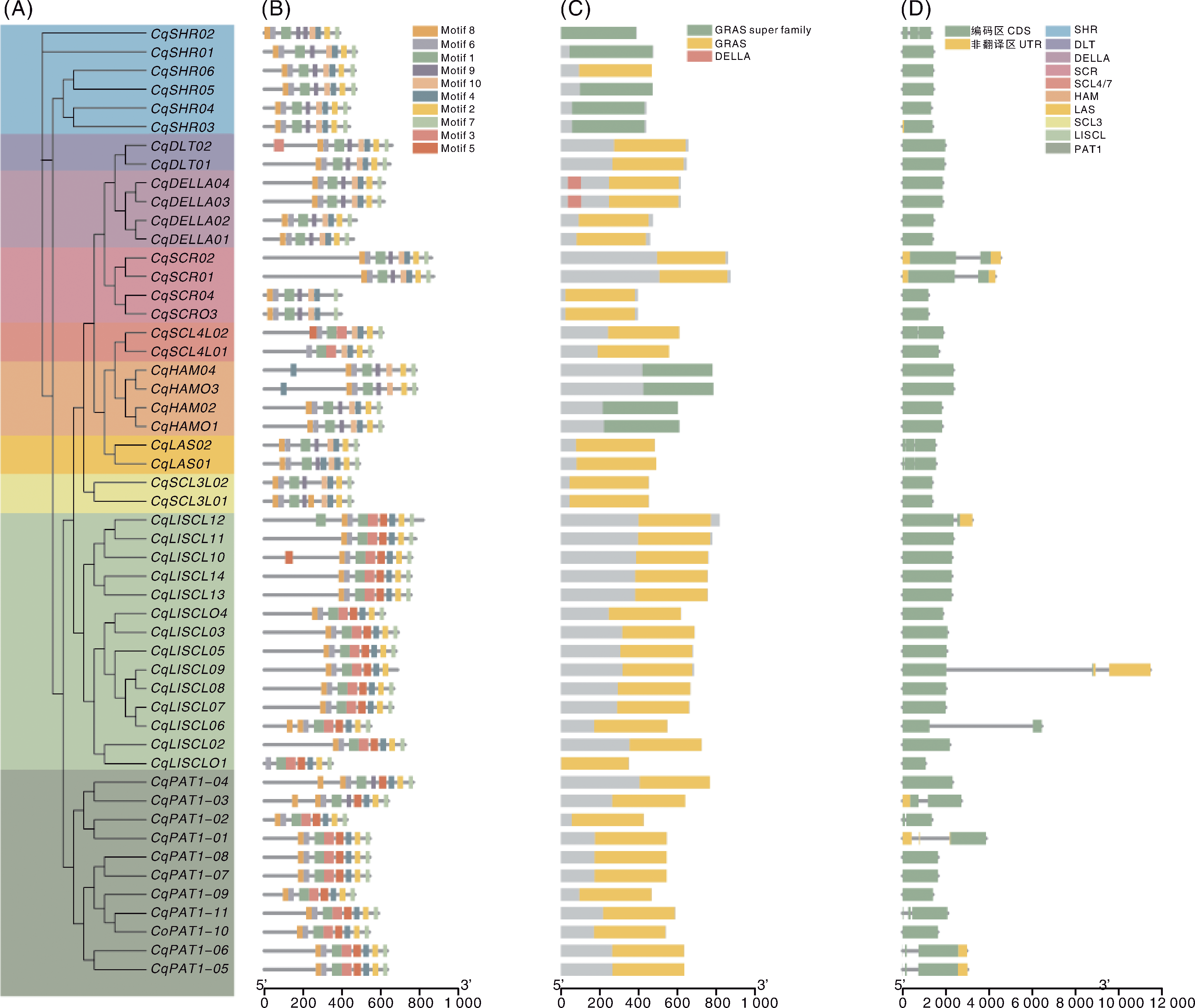
Fig.2 Clustering, conserved motifs, conserved domains, and gene structure composition of quinoa GRAS gene family members A, Clustering of quinoa GRAS gene family members (different colored regions represent different subfamilies); B, Composition and distribution of conserved motifs of CqGRAS; C, Conserved domains of the CqGRAS family proteins; D. Gene structure of CqGRAS.
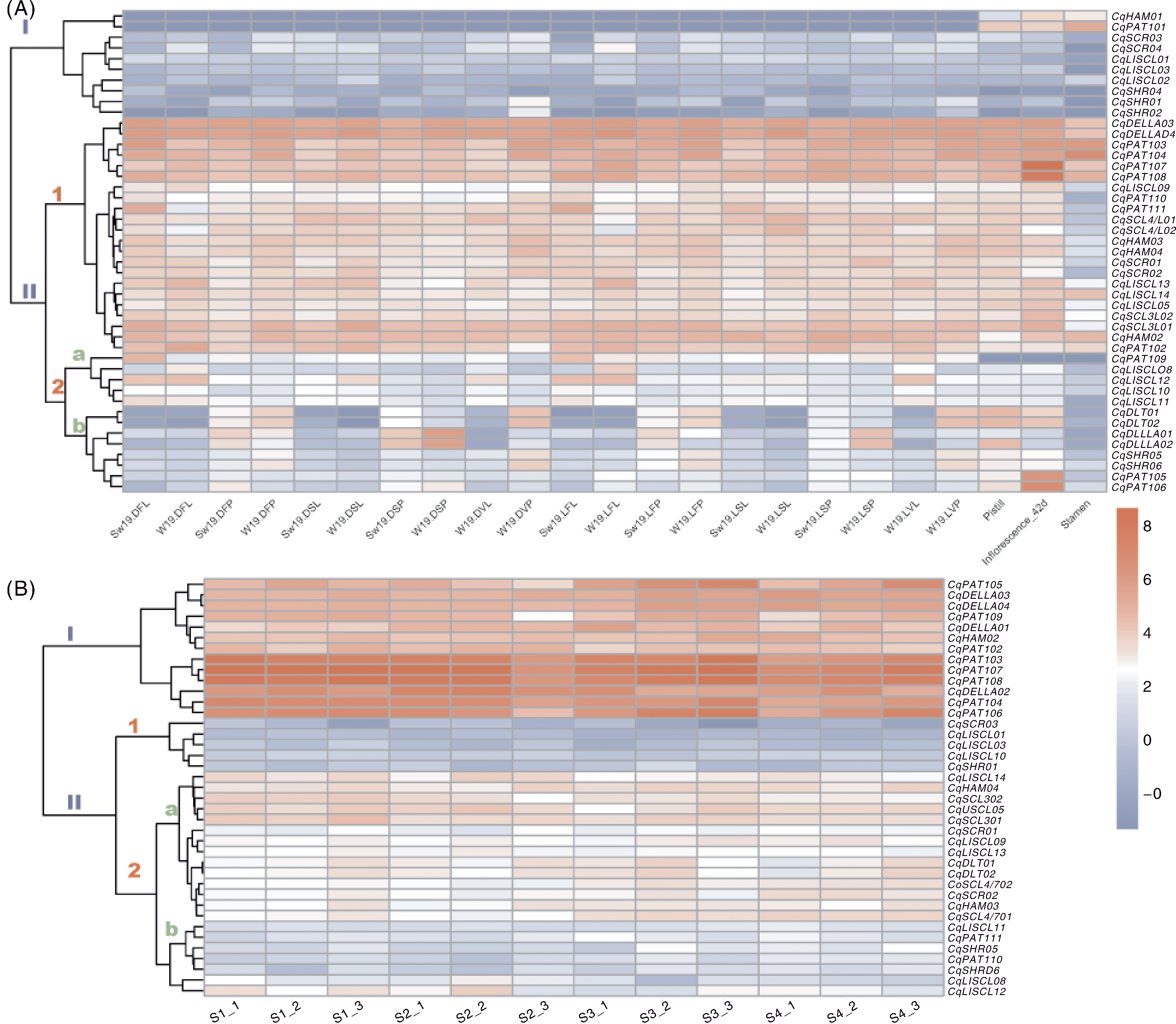
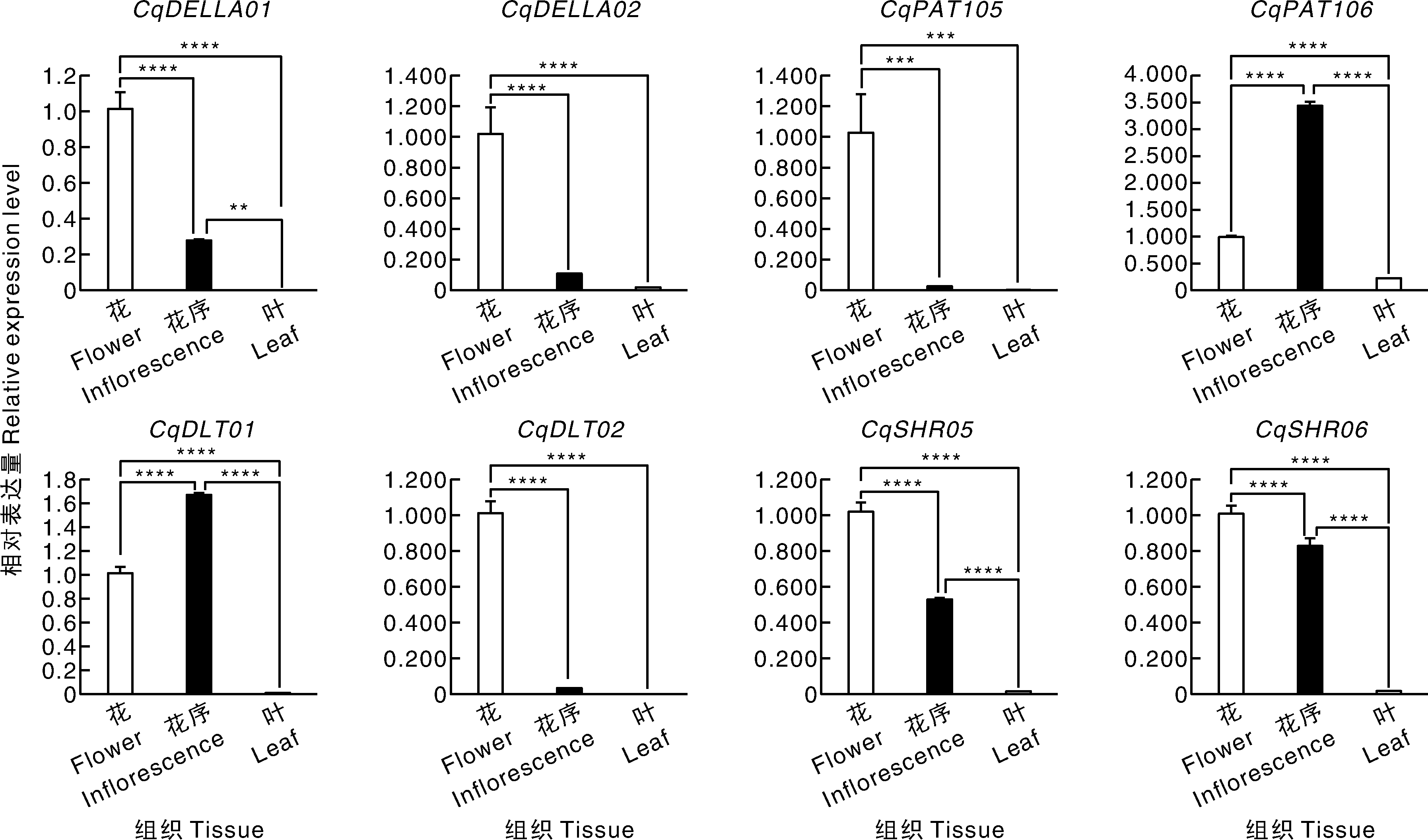
Fig.4 Expression of GRAS genes in quinoa at different developmental stages and seed development A, Expression of GRAS genes in leaves, inflorescences, stamens, and pistils at different developmental stages. Sample naming: S, Summer; W, Winter; 19, W19 variety; D, Short-day (14 h light/10 h dark) condition; L, Long-day (22 h light/2 h dark) condition; V, Vegetative stage; F, Flowering stage; S, Seed formation stage; P, Panicle sample; L, Leaf sample. e.g., S19DFL represented summer W19 flowering stage leaves under short-day conditions. B, Expression of GRAS genes during seed development. S1-S4 represent the 7 d (S1), 17 d (S2), 27 d (S3), and 42 d (S4) after flowering in the TY-20, with three biological replicates (1, 2, 3).
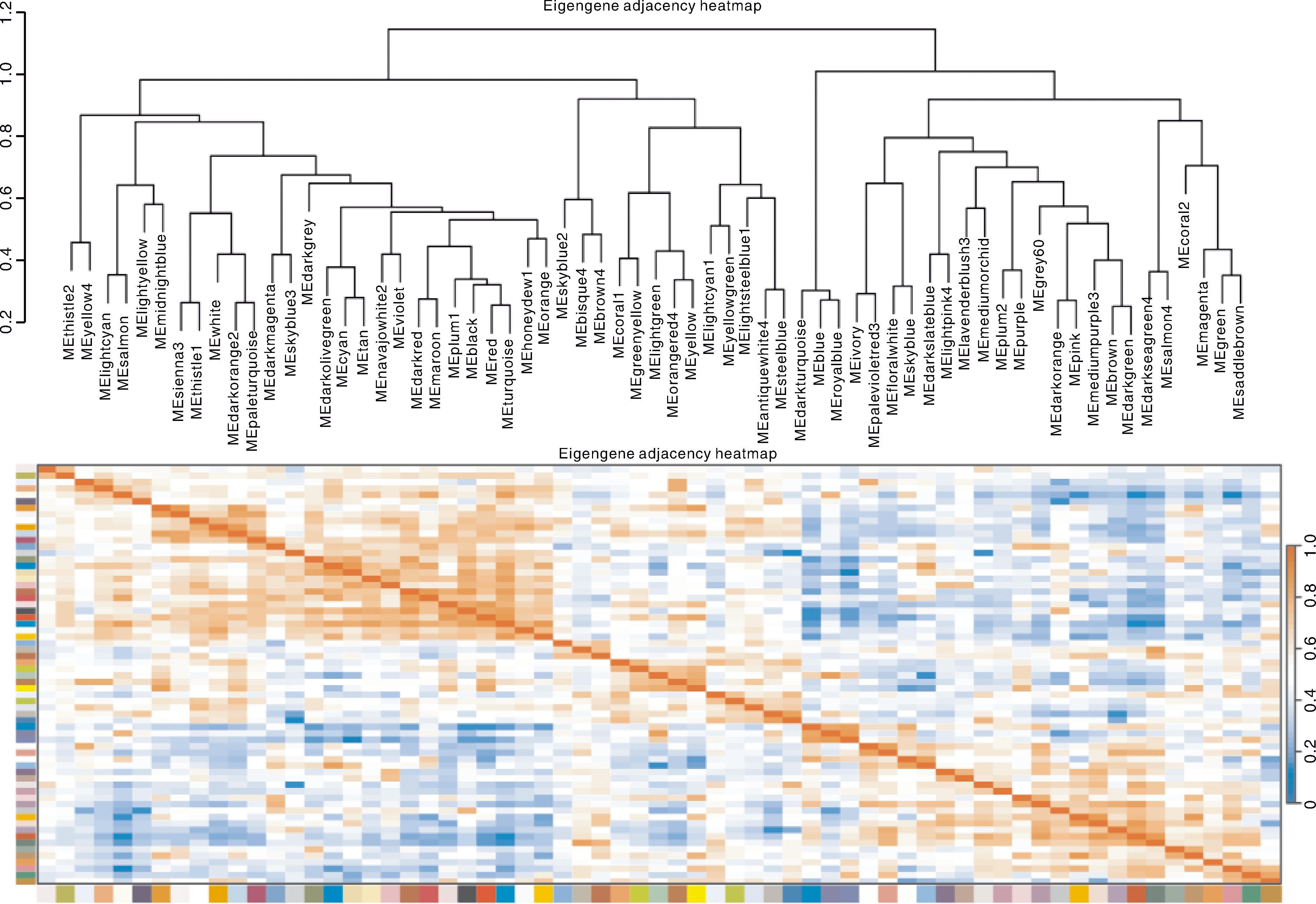
Fig.6 Clustering and heat map between modules Red color of each box represents the positive correlation between modules. Blue color of each box represents the negative correlation between modules.
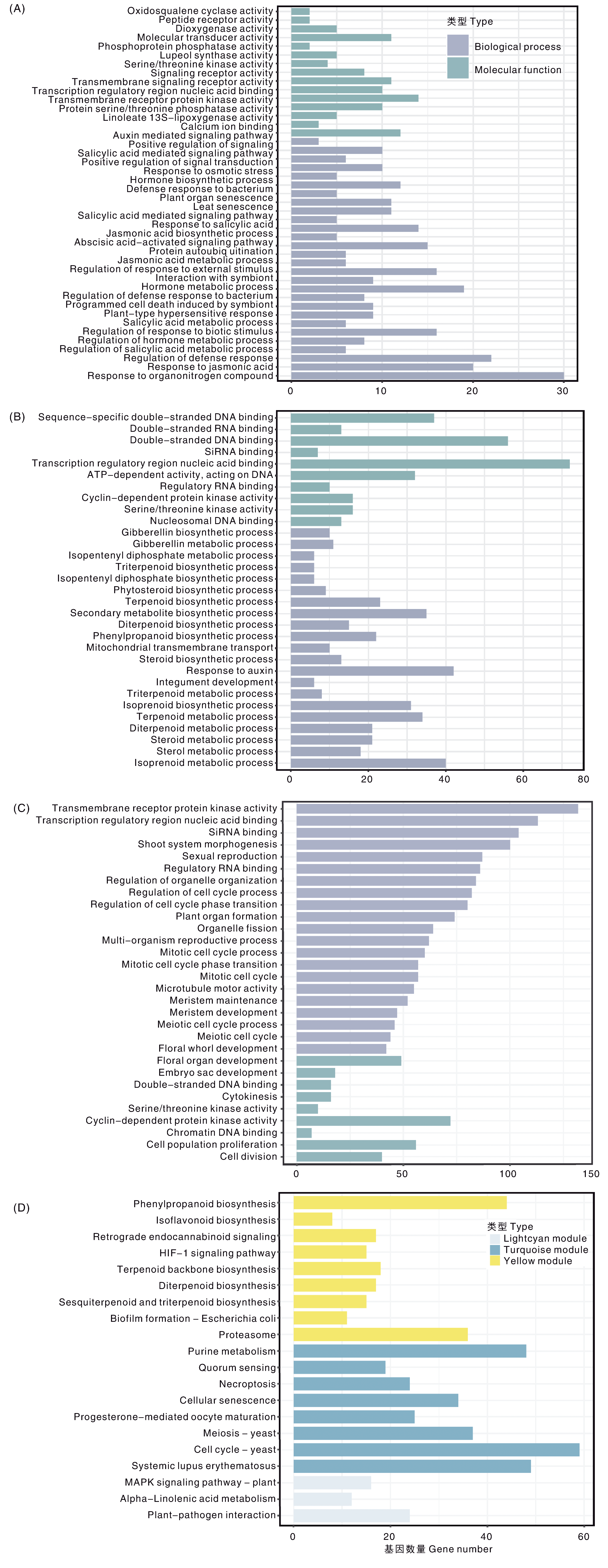
Fig.7 GO enrichment and KEGG enrichment of module genes in quinoa A-C, GO enrichment module of lightcyan, yellow and turquoise, respectively; D, KEGG enrichment of three combining module genes.
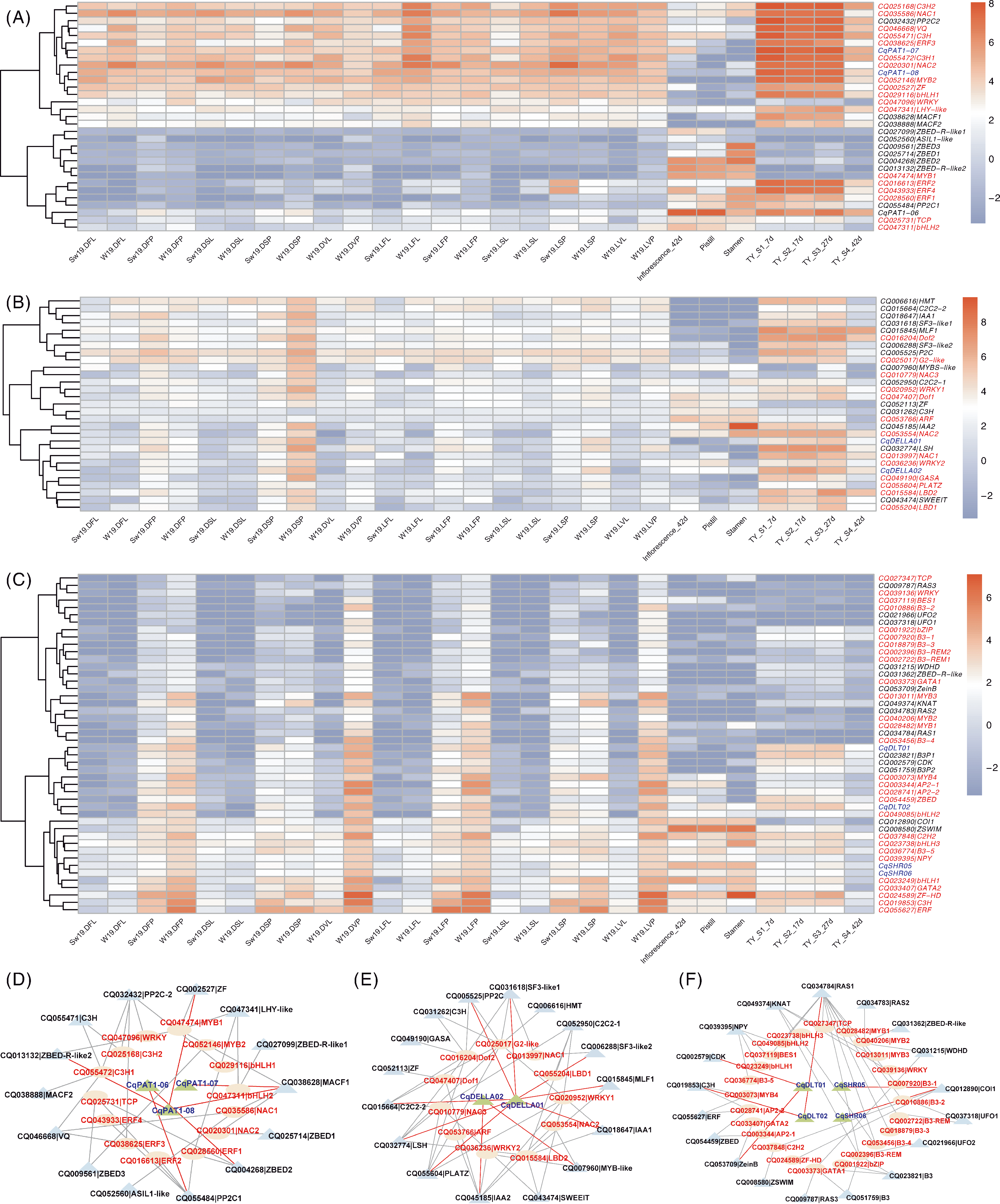
Fig.8 Heat map and correlation of gene and transcription factor expression in flower development related tissues within the module A-C represented the expression of CqGRAS genes and transcription factors in flower and seed development related tissues in the lightcyan,yellow and tuequoise modules; D-F represented the network diagram of CqGRAS genes and related transcription factors in the lightcyan,yellow and tuequoise modules. Green triangles represented CqGRAS members, blue triangles represented other genes that interact with transcription factors, and ellipses represented transcription factors.
| [1] | PENG J R, CAROL P, RICHARDS D E, et al. The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses[J]. Genes & Development, 1997, 11(23): 3194-3205. |
| [2] | SILVERSTONE A L, CIAMPAGLIO C N, SUN T P. The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway[J]. The Plant Cell, 1998, 10(2): 155-169. |
| [3] | SAMACH A, KLENZ J E, KOHALMI S E, et al. The unusual floral organs gene of Arabidopsis thaliana is an F-box protein required for normal patterning and growth in the floral meristem[J]. The Plant Journal, 1999, 20(4): 433-445. |
| [4] | BOLLE C. The role of GRAS proteins in plant signal transduction and development[J]. Planta, 2004, 218(5): 683-692. |
| [5] | HIRANO Y, NAKAGAWA M, SUYAMA T, et al. Structure of the SHR-SCR heterodimer bound to the BIRD/IDD transcriptional factor JKD[J]. Nature Plants, 2017, 3: 17010. |
| [6] | PARK M J, KWON Y J, GIL K E, et al. LATE ELONGATED HYPOCOTYL regulates photoperiodic flowering via the circadian clock in Arabidopsis[J]. BMC Plant Biology, 2016, 16(1): 114. |
| [7] | SUN T P, GUBLER F. Molecular mechanism of gibberellin signaling in plants[J]. Annual Review of Plant Biology, 2004, 55: 197-223. |
| [8] | CENCI A, ROUARD M. Evolutionary analyses of GRAS transcription factors in angiosperms[J]. Frontiers in Plant Science, 2017, 8: 273. |
| [9] | MOROHASHI K, MINAMI M, TAKASE H, et al. Isolation and characterization of a novel GRAS gene that regulates meiosis-associated gene expression[J]. Journal of Biological Chemistry, 2003, 278(23): 20865-20873. |
| [10] | SABATINI S, HEIDSTRA R, WILDWATER M, et al. SCARECROW is involved in positioning the stem cell niche in the Arabidopsis root meristem[J]. Genes & Development, 2003, 17(3): 354-358. |
| [11] | EDGAR R C. MUSCLE: multiple sequence alignment with high accuracy and high throughput[J]. Nucleic Acids Research, 2004, 32(5): 1792-1797. |
| [12] | TONG H N, JIN Y, LIU W B, et al. DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice[J]. The Plant Journal, 2009, 58(5): 803-816. |
| [13] | ZHU X L, WANG B Q, WEI X H. Genome wide identification and expression pattern analysis of the GRAS family in quinoa[J]. Functional Plant Biology, 2021, 48(9): 948-962. |
| [14] | LIU X Y, WIDMER A. Genome-wide comparative analysis of the GRAS gene family in Populus, Arabidopsis and rice[J]. Plant Molecular Biology Reporter, 2014, 32(6): 1129-1145. |
| [15] | LING H, ZENG X, GUO S X. Functional insights into the late embryogenesis abundant (LEA) protein family from Dendrobium officinale(Orchidaceae) using an Escherichia coli system[J]. Scientific Reports, 2016, 6: 39693. |
| [16] | MCGINNIS S, MADDEN T L. BLAST: at the core of a powerful and diverse set of sequence analysis tools[J]. Nucleic Acids Research, 2004, 32: W20-W25. |
| [17] | DUVAUD S, GABELLA C, LISACEK F, et al. Expasy, the Swiss bioinformatics resource portal, as designed by its users[J]. Nucleic Acids Research, 2021, 49(W1): W216-W227. |
| [18] | HORTON P, PARK K J, OBAYASHI T, et al. WoLF PSORT: protein localization predictor[J]. Nucleic Acids Research, 2007, 35: W585-W587. |
| [19] | DREWS G N, BOWMAN J L, MEYEROWITZ E M. Negative regulation of the Arabidopsis homeotic gene AGAMOUS by the APETALA2 product[J]. Cell, 1991, 65(6): 991-1002. |
| [20] | RUEDA-ROMERO P, BARRERO-SICILIA C, GÓMEZ-CADENAS A, et al. Arabidopsis thaliana DOF6 negatively affects germination in non-after-ripened seeds and interacts with TCP14[J]. Journal of Experimental Botany, 2012, 63(5): 1937-1949. |
| [21] | URBANOVA T, LEUBNER-METZGER G. Gibberellins and seed germination[M]// HEDDEN P, THOMAS S G. Annual plant reviews. Hoboken: John Wiley & Sons, Inc., 2016: 253-284. |
| [22] | LETUNIC I, BORK P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees[J]. Nucleic Acids Research, 2016, 44(W1): W242-W245. |
| [23] | WANG Y P, TANG H B, DEBARRY J D, et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity[J]. Nucleic Acids Research, 2012. 40(7): e49. |
| [24] | CHEN C J, WU Y, XIA R. A painless way to customize Circos plot: from data preparation to visualization using TBtools[J]. iMeta, 2022, 1(3): e35. |
| [25] | HU B, JIN J P, GUO A Y, et al. GSDS 2.0: an upgraded gene feature visualization server[J]. Bioinformatics, 2015, 31(8): 1296-1297. |
| [26] | BAILEY T L, JOHNSON J, GRANT C E, et al. The MEME suite[J]. Nucleic Acids Research, 2015, 43(W1): W39-W49. |
| [27] | MARCHLER-BAUER A, DERBYSHIRE M K, GONZALES N R, et al. CDD: NCBI’s conserved domain database[J]. Nucleic Acids Research, 2015, 43(D1): D222-D226. |
| [28] | CHEN C J, CHEN H, ZHANG Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data[J]. Molecular Plant, 2020, 13(8): 1194-1202. |
| [29] | SHEN W, SIPOS B, ZHAO L Y. SeqKit2: a Swiss army knife for sequence and alignment processing[J]. iMeta, 2024, 3(3): e191. |
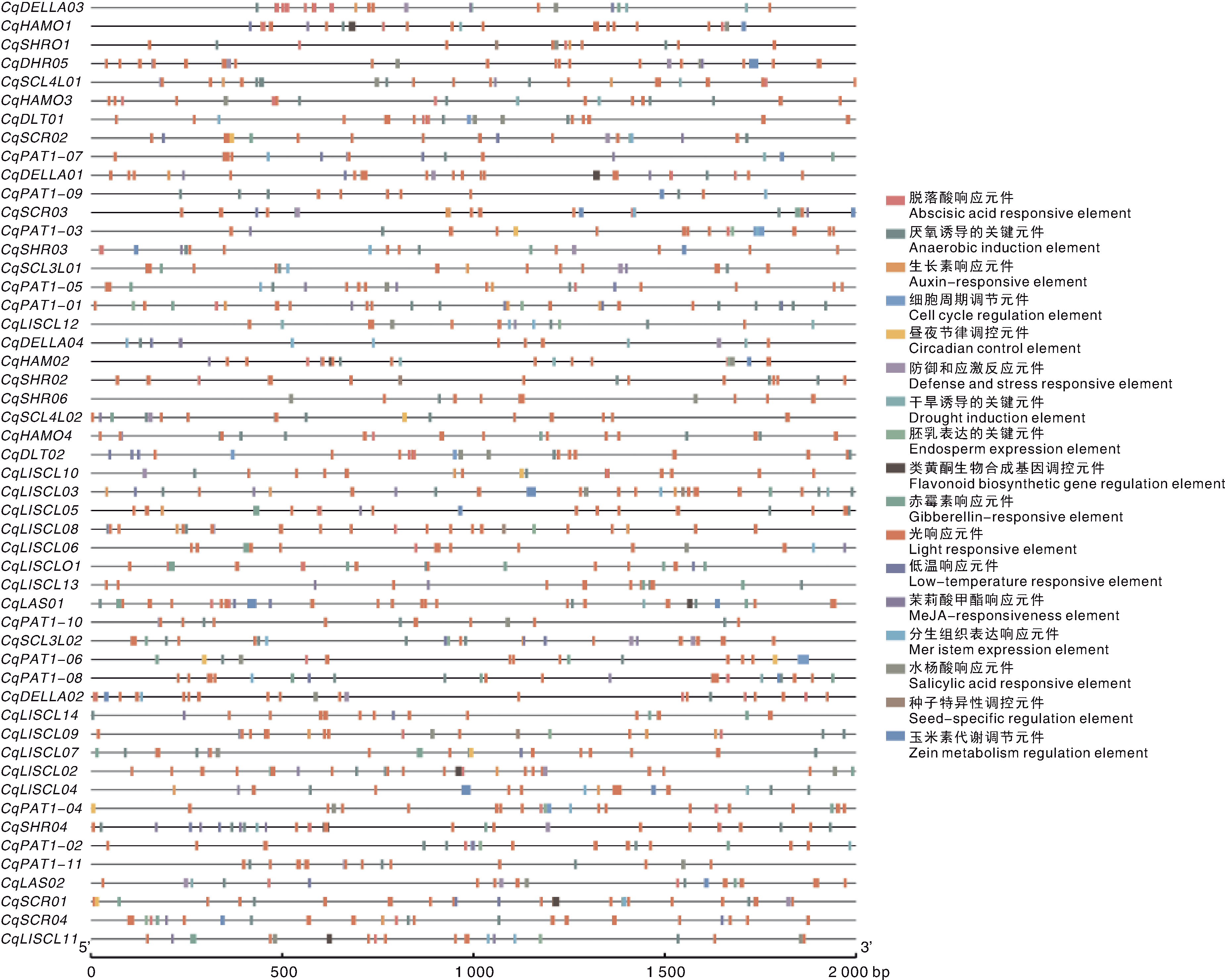
| [30] | LESCOT M. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences[J]. Nucleic Acids Research, 2002, 30(1): 325-327. |
| [31] | SHAN Z Y, LUO X L, WU M Y, et al. Genome-wide identification and expression of GRAS gene family members in cassava[J]. BMC Plant Biology, 2020, 20(1): 46. |
| [32] | CHEN S F. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp[J]. iMeta, 2023, 2(2): e107. |
| [33] | KIM D, PAGGI J M, PARK C, et al. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype[J]. Nature Biotechnology, 2019, 37(8): 907-915. |
| [34] | DANECEK P, BONFIELD J K, LIDDLE J, et al. Twelve years of SAMtools and BCFtools[J]. GigaScience, 2021, 10(2): giab008. |
| [35] | LIAO Y, SMYTH G K, SHI W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features[J]. Bioinformatics, 2014, 30(7): 923-930. |
| [36] | DAILEY A L. Metabolomic bioinformatic analysis[M]// Methods in molecular biology. New York: Springer New York, 2017: 341-352. |
| [37] | LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔT method[J]. Methods, 2001, 25(4): 402-408. |
| [38] | LANGFELDER P, HORVATH S. WGCNA: an R package for weighted correlation network analysis[J]. BMC Bioinformatics, 2008, 9(1): 559. |
| [39] | WU T Z, HU E Q, XU S B, et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data[J]. The Innovation, 2021, 2(3): 100141. |
| [40] | SHANNON P, MARKIEL A, OZIER O, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks[J]. Genome Research, 2003, 13(11): 2498-2504. |
| [41] | LEE K C, JANG Y H, KIM S K, et al. RRM domain of Arabidopsis splicing factor SF1 is important for pre-mRNA splicing of a specific set of genes[J]. Plant Cell Reports, 2017, 36(7): 1083-1095. |
| [42] | WILLIAMS J H, DALY L N, INGLEY E, et al. HLS7 a hemopoietic lineage switch gene homologous to the leukemia-inducing gene MLF1[J]. The EMBO Journal, 1999, 18(20): 5559-5566. |
| [43] | CHO E, ZAMBRYSKI P C. ORGAN BOUNDARY1 defines a gene expressed at the junction between the shoot apical meristem and lateral organs[J]. Proceedings of the National Academy of Sciences of the United States of America, 2011, 108(5): 2154-2159. |
| [44] | CHEN Y Q, TAI S S, WANG D W, et al. Homology-based analysis of the GRAS gene family in tobacco[J]. Genetics and Molecular Research, 2015, 14(4): 15188-15200. |
| [45] | SONG X M, LIU T K, DUAN W K, et al. Genome-wide analysis of the GRAS gene family in Chinese cabbage (Brassica rapa ssp. pekinensis)[J]. Genomics, 2014, 103(1): 135-146. |
| [46] | STROUD M J, NAZGIEWICZ A, MCKENZIE E A, et al. GAS2-like proteins mediate communication between microtubules and actin through interaction with end-binding proteins[J]. Journal of Cell Science, 2014: jcs.140558. |
| [47] | ZHANG H, LIU X Q, WANG X M, et al. Genome-wide identification of GRAS gene family and their responses to abiotic stress in Medicago sativa[J]. International Journal of Molecular Sciences, 2021, 22(14): 7729. |
| [48] | GAZZARRINI S, LEJAY L, GOJON A, et al. Three functional transporters for constitutive, diurnally regulated, and starvation-induced uptake of ammonium into Arabidopsis roots[J]. The Plant Cell, 1999, 11(5): 937. |
| [49] | MOURADOV A, CREMER F, COUPLAND G. Control of flowering time: interacting pathways as a basis for diversity[J]. The Plant Cell, 2002, 14(suppl 1): S111-S130. |
| [50] | MOON J, LEE H, KIM M, et al. Analysis of flowering pathway integrators in Arabidopsis[J]. Plant & Cell Physiology, 2005, 46(2): 292-299. |
| [51] | ZHAO H, BAO Y. PIF4: integrator of light and temperature cues in plant growth[J]. Plant Science, 2021, 313: 111086. |
| [52] | BUNDOCK P, HOOYKAAS P. An Arabidopsis hAT-like transposase is essential for plant development[J]. Nature, 2005, 436(7048): 282-284. |
| [53] | CHEN S S, DENG J R, CHENG P D, et al. Transcriptome-wide identification of walnut PP2C family genes in response to external stimulus[J]. BMC Genomics, 2022, 23(1): 640. |
| [54] | DAVIÈRE J M, ACHARD P. Gibberellin signaling in plants[J]. Development, 2013, 140(6): 1147-1151. |
| [55] | MAI Y X, WANG L, YANG H Q. A gain-of-function mutation in IAA7/AXR2 confers late flowering under short-day light in Arabidopsis[J]. Journal of Integrative Plant Biology, 2011, 53(6): 480-492. |
| [56] | RIEU I, RUIZ-RIVERO O, FERNANDEZ-GARCIA N, et al. The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle[J]. The Plant Journal, 2008, 53(3): 488-504. |
| [57] | ZHANG S C, YANG C W, PENG J Z, et al. GASA5, a regulator of flowering time and stem growth in Arabidopsis thaliana[J]. Plant Molecular Biology, 2009, 69(6): 745-759. |
| [58] | ZHANG Y Q, LIU Z J, LIU J P, et al. GA-DELLA pathway is involved in regulation of nitrogen deficiency-induced anthocyanin accumulation[J]. Plant Cell Reports, 2017, 36(4): 557-569. |
| [59] | MÉNDEZ-BRAVO A, RUIZ-HERRERA L F, CRUZ-RAMÍREZ A, et al. CONSTITUTIVE TRIPLE RESPONSE1 and PIN2 act in a coordinate manner to support the indeterminate root growth and meristem cell proliferating activity in Arabidopsis seedlings[J]. Plant Science, 2019, 280: 175-186. |
| [60] | XU L H, LIU F Q, LECHNER E, et al. The SCF(COI1) ubiquitin-ligase complexes are required for jasmonate response in Arabidopsis[J]. The Plant Cell, 2002, 14(8): 1919-1935. |
| [61] | TANG Y X, GAO X, CUI Y N, et al. Research advances in the plant TCP transcription factors[J]. Chinese Science Bulletin, 2022, 67(33): 3964-3975. |
| [62] | KEN H G, TAKANO M, NEUMANN R, et al. The rice COLEOPTILE PHOTOTROPISM1 gene encoding an ortholog of Arabidopsis NPH3 is required for phototropism of coleoptiles and lateral translocation of auxin[J]. The Plant Cell, 2005, 17(1): 103-115. |
| [63] | LEVIN J Z, MEYEROWITZ E M. UFO: an Arabidopsis gene involved in both floral meristem and floral organ development[J]. The Plant Cell, 1995, 7(5): 529. |
| [64] | KROGAN N T, HOGAN K, LONG J A. APETALA2 negatively regulates multiple floral organ identity genes in Arabidopsis by recruiting the co-repressor TOPLESS and the histone deacetylase HDA19[J]. Development, 2012, 139(22): 4180-4190. |
| [65] | PRESS M O, LANCTOT A, QUEITSCH C. PIF4 and ELF3 act independently in Arabidopsis thaliana thermoresponsive flowering[J]. PLoS One, 2016, 11(8): e0161791. |
| [66] | DALL’OSTO L, CAZZANIGA S, NORTH H, et al. The Arabidopsis Aba4-1 mutant reveals a specific function for neoxanthin in protection against photooxidative stress[J]. The Plant Cell, 2007, 19(3): 1048-1064. |
| [67] | LI X Y, QIAN Q, FU Z M, et al. Control of tillering in rice[J]. Nature, 2003, 422(6932): 618-621. |
| [68] | WANG L, SUN S Y, JIN J Y, et al. Coordinated regulation of vegetative and reproductive branching in rice[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(50): 15504-15509. |
| [69] | VARSHNEY R K, GRANER A, SORRELLS M E. Genomics-assisted breeding for crop improvement[J]. Trends in Plant Science, 2005, 10(12): 621-630. |
| [70] | PEARCE S, SAVILLE R, VAUGHAN S P, et al. Molecular characterization of Rht-1 dwarfing genes in hexaploid wheat[J]. Plant Physiology, 2011, 157(4): 1820-1831. |
| [71] | COLLARD B C Y, MACKILL D J. Marker-assisted selection: an approach for precision plant breeding in the twenty-first century[J]. Philosophical Transactions of the Royal Society B: Biological Sciences, 2008, 363(1491): 557-572. |
| [72] | JARVIS D E, HO Y S, LIGHTFOOT D J, et al. The genome of Chenopodium quinoa[J]. Nature, 2017, 542(7641): 307-312. |
| [73] | RUIZ K B, BIONDI S, MARTÍNEZ E A, et al. Quinoa: a model crop for understanding salt-tolerance mechanisms in halophytes[J]. Plant Biosystems-an International Journal Dealing with All Aspects of Plant Biology, 2016, 150(2): 357-371. |
| [74] | TESTER M, LANGRIDGE P. Breeding technologies to increase crop production in a changing world[J]. Science, 2010, 327(5967): 818-822. |
| [75] | OGATA T, TOYOSHIMA M, YAMAMIZO-ODA C, et al. Virus-mediated transient expression techniques enable functional genomics studies and modulations of betalain biosynthesis and plant height in quinoa[J]. Frontiers in Plant Science, 2021, 12: 643499. |
| [1] | ZHENG Ting, XIANG Jiang, WEI Lingzhu, WU Jiang, CHENG Jianhui. Analysis on the effects of CPPU and TDZ on the aroma of Tiangong Moyu grape using WGCNA and the exploration of the key genes [J]. Acta Agriculturae Zhejiangensis, 2025, 37(2): 311-320. |
| [2] | WU Tao, WEI Yuming, JIANG Xiaofan, HUANG Jie, YANG Farong, CHEN Guoshun, CAI Yuan, JIAO Ting, ZHAO Shenguo. Effects of Chenopodium quinoa in diets on growth performance, slaughter performance, organ index and intestinal morphology of Luhua chickens [J]. Acta Agriculturae Zhejiangensis, 2022, 34(2): 255-265. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||