浙江农业学报 ›› 2026, Vol. 38 ›› Issue (2): 405-416.DOI: 10.3969/j.issn.1004-1524.20250179
• 综述 • 上一篇
裂解多糖单加氧酶的生物学机制及其在木质纤维素降解中的应用
朱夏菁1,2( ), 孙宏1, 周航海1, 王新1, 皮二旭2,*(
), 孙宏1, 周航海1, 王新1, 皮二旭2,*( ), 汤江武1,3,*(
), 汤江武1,3,*( )
)
1.浙江省农业科学院 环境资源与土壤肥料研究所 浙江 杭州 310021 2.杭州师范大学 生命与环境科学学院 浙江 杭州 311121 3.浙江省农业科学院 园艺研究所 浙江 杭州 310021
-
收稿日期:2025-03-12出版日期:2026-02-25发布日期:2026-03-24 -
作者简介:朱夏菁,研究方向为真菌来源LPMO降解难分解天然生物质。E-mail:zhuxjchn@163.com -
通讯作者:*皮二旭,E-mail:erxupi@hznu.edu.cn;
汤江武,E-mail:angjiangwu@sina.com -
基金资助:浙江省自然科学基金杭州区域创新发展联合基金(LHZSZ24C010001);浙江省农业科学院院地科技合作项目(ZJTY2024-A-55)
Biological mechanism of lytic polysaccharide monooxygenase and its applications in lignocellulose degradation
ZHU Xiajing1,2( ), SUN Hong1, ZHOU Hanghai1, WANG Xin1, PI Erxu2,*(
), SUN Hong1, ZHOU Hanghai1, WANG Xin1, PI Erxu2,*( ), TANG Jiangwu1,3,*(
), TANG Jiangwu1,3,*( )
)
1. Institute of Environmental Resources ,Soil and Fertilizer, Zhejiang Academy of Agricultural Sciences Hangzhou 310021, China 2. College of Life and Environmental Sciences ,Hangzhou Normal University Hangzhou 311121, China 3. Institute of Horticulture ,Zhejiang Academy of Agricultural Sciences Hangzhou 310021, China
-
Received:2025-03-12Published:2026-02-25Online:2026-03-24
摘要:
裂解多糖单加氧酶(LPMO)是一种氧化酶,其生化特性、在木质纤维素降解中的作用、蛋白质工程改造与优化,以及在工业生物技术中的应用受到广泛关注。木质纤维素因其复杂的结构与化学成分在天然条件下难以高效降解,其有效降解与转化已成为一项重要挑战。LPMO能够通过氧化作用切断多糖的结晶表面,协同糖苷水解酶高效降解木质纤维素,从而在秸秆等天然难降解生物质的转化中发挥重要作用。采用异源表达在其他宿主中生产LPMO是一种提升LPMO产量并推动其产业化的有效策略,当前已有诸多成功案例通过基因工程手段提升LPMO的稳定性、活性、底物特异性与催化效率。目前,LPMO已在植物抗病、食品工业与畜牧工业等工业生物技术领域展现出巨大的应用潜力。未来研究可进一步深入解析LPMO的催化降解机制,拓展其高效异源表达与蛋白质工程改造策略,以推动LPMO的工业化应用。
中图分类号:
引用本文
朱夏菁, 孙宏, 周航海, 王新, 皮二旭, 汤江武. 裂解多糖单加氧酶的生物学机制及其在木质纤维素降解中的应用[J]. 浙江农业学报, 2026, 38(2): 405-416.
ZHU Xiajing, SUN Hong, ZHOU Hanghai, WANG Xin, PI Erxu, TANG Jiangwu. Biological mechanism of lytic polysaccharide monooxygenase and its applications in lignocellulose degradation[J]. Acta Agriculturae Zhejiangensis, 2026, 38(2): 405-416.
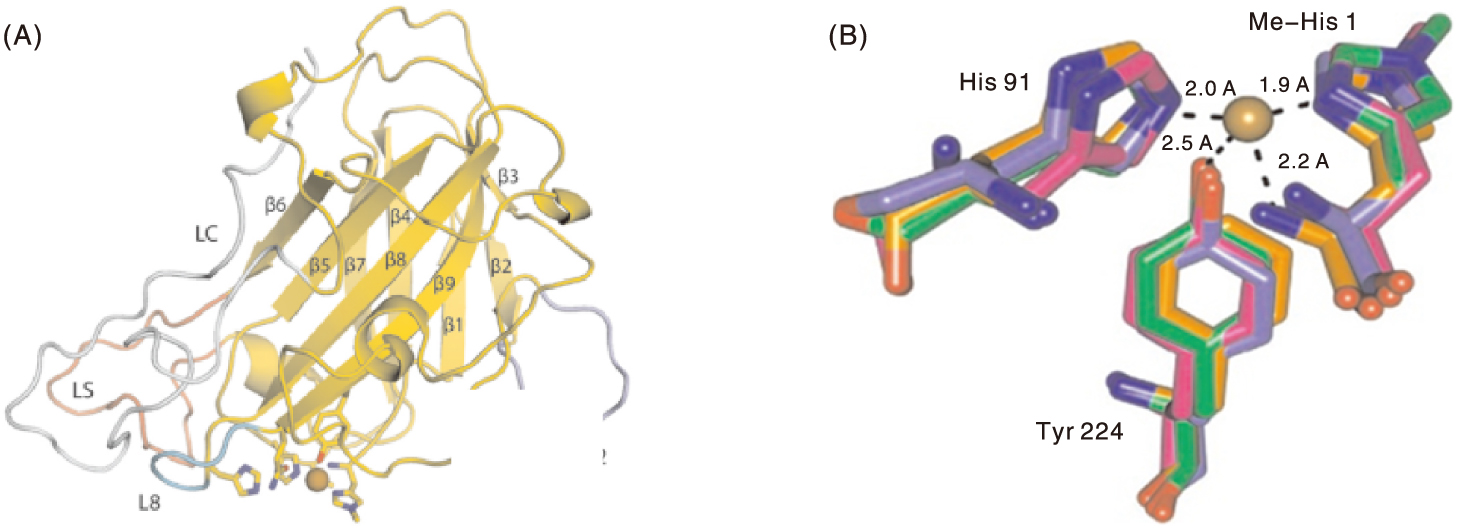
图1 LPMO及其活性中心的三维结构 A,LPMO三维结构示意图;B,4个家族LPMO活性中心三维结构叠加示意图。棕色球体,铜原子;洋红色结构,AA9(PDB:3ZUD);橙色结构,AA10(PDB:2YOY);紫色结构,AA11(PDB:4MAI);绿色结构,AA13(PDB:4OPB)。His,组氨酸;Tyr,酪氨酸;Me-His 1,甲基化组氨酸。
Fig.1 Typical three-dimensional structure of LPMO and its active center A, Three-dimensional structure schematic of LPMO; B, Superposition of three-dimensional structures of active centers from four LPMO families. Brown sphere, Copper atom; magenta structure, AA9 (PDB:3ZUD); Orange structure, AA10 (PDB:2YOY); Purple structure, AA11 (PDB:4MAI); Green structure, AA13 (PDB:4OPB). His, Histidine; Tyr, Tyrosine; Me-His 1, Methylated histidine.
| LPMO名称 LPMO name | 表达系统 Expression system | 参考文献 Reference |
|---|---|---|
| PcLPMO9D、NcLPMO9C、MtLPMO9E | 毕赤酵母(pPICZαA) P. pastoris(pPICZαA) | [ |
| SmLPMO10A、BlLPMO10A、JdLPMO10A、 ScLPMO10C、MaLPMO10B、CjLPMO10A | 大肠埃希菌(基于XylS/Pm系统的新型LPMO表达克隆载体) E. coli(novel LPMO expression cloning vector based on the XylS/Pm system) | [ |
| CjLPMO10A | 大肠埃希菌(pET22b) E. coli(pET22b) | [ |
| 米曲霉AA11家族LPMO A. oryzae AA11 family LPMO | 大肠埃希菌(pET32a)和毕赤酵母(pPIC9K) E. coli(pET32a) and P. pastoris(pPIC9K) | [ |
| TfLPMO10A、ScLPMO10B、 BatLPMO10、LsAA9A | 大肠埃希菌(pLyGo-Ec-1、pLyGo-Ec-2、pLyGo-Ec-3等多种载体) E. coli(various vectors including pLyGo-Ec-1, pLyGo-Ec-2, pLyGo-Ec-3) | [ |
表1 LPMO的基因工程表达系统
Table 1 Expression system of LPMO genetic engineering
| LPMO名称 LPMO name | 表达系统 Expression system | 参考文献 Reference |
|---|---|---|
| PcLPMO9D、NcLPMO9C、MtLPMO9E | 毕赤酵母(pPICZαA) P. pastoris(pPICZαA) | [ |
| SmLPMO10A、BlLPMO10A、JdLPMO10A、 ScLPMO10C、MaLPMO10B、CjLPMO10A | 大肠埃希菌(基于XylS/Pm系统的新型LPMO表达克隆载体) E. coli(novel LPMO expression cloning vector based on the XylS/Pm system) | [ |
| CjLPMO10A | 大肠埃希菌(pET22b) E. coli(pET22b) | [ |
| 米曲霉AA11家族LPMO A. oryzae AA11 family LPMO | 大肠埃希菌(pET32a)和毕赤酵母(pPIC9K) E. coli(pET32a) and P. pastoris(pPIC9K) | [ |
| TfLPMO10A、ScLPMO10B、 BatLPMO10、LsAA9A | 大肠埃希菌(pLyGo-Ec-1、pLyGo-Ec-2、pLyGo-Ec-3等多种载体) E. coli(various vectors including pLyGo-Ec-1, pLyGo-Ec-2, pLyGo-Ec-3) | [ |
| 改造内容 Modification content | 具体结果 Specific result | 参考文献 Reference |
|---|---|---|
| 优化密码子和替换同义密码子, 对LPMO基因进行优化,通过优 化培养条件提高酶表达量,增强 酶活性 Codon optimization and synonymous codon replacement for LPMO gene, enhanced enzyme expression and activity through optimized culture conditions | 优化培养时间、诱导温度与甲醇添加量后,重组毕赤酵母菌株产LPMO能力较优化前增加了12.51%,重组菌株的酶活性最高达到(25.64±0.21) U·L-1;以1%微晶纤维素为底物时,添加LPMO可以使酶解6 h后水解率提升26.46%;以汽爆玉米秸秆为底物时,添加LPMO和一定浓度抗坏血酸可以缩短纤维素酶的降解启动时间,并在酶解9 h后水解效率提升13.07% After optimizing culture time, induction temperature, and methanol addition, the LPMO production of recombinant P. pastoris increased by 12.51% compared with that before optimization. The maximum enzyme activity reached (25.64±0.21) U·L-1. When 1% microcrystalline cellulose was used as the substrate, the hydrolysis rate increased by 26.46% after 6 h of enzymatic hydrolysis with LPMO. When steam-exploded corn stover was used as the substrate, the addition of LPMO and a certain concentration of ascorbic acid shortened the degradation initiation time of cellulase and increased the hydrolysis efficiency by 13.07% after 9 h of enzymatic hydrolysis | [ |
| 构建含有不同信号肽的表达载体, 提高酶表达量 Construction of expression vectors with different signal peptides to enhance enzyme expression | 使用结合了XylS启动子/激活子系统和不同LPMO信号序列的载体,该新型表达系统能可靠产生成熟的LPMO,并且使用SmLPMO10A的信号肽时受试LPMO的表达水平显著提高,SmLPMO10A、BlLPMO10A、JdLPMO10A的产量分别达到22、7、10 mg·L-1 Using a carrier combining the XylS promoter/activator system and different LPMO signal sequences, this novel expression system reliably produces mature LPMO. The expression levels of the tested LPMOs were significantly increased when using the signal peptide of SmLPMO10A, with yields of SmLPMO10A, BlLPMO10A, and JdLPMO10A reaching 22, 7, and 10 mg·L-1, respectively | [ |
| 引入二硫键,提高热稳定性、化学 稳定性和活性 The disulfide bond was introduced to improve the thermal stability, chemical stability and activity | 与野生型相比,突变体在60 ℃下的半衰期增加3倍,表观Tm值上升7 ℃,化学变性中点浓度(midpoint concentration of chemical denaturation, Cm值)提高0.3 mol·L-1,显示出野生型1.5倍的酶活性 Compared with the wild type, the mutant had a 3-fold increased half-life at 60 ℃, a 7 ℃ increase in apparent Tm value, a 0.3 mol·L-1 increase in midpoint concentration of chemical denaturation(Cm value), and 1.5 times the enzyme activity of the wild type | [ |
| 异源表达,提高酶表达量和催化 活性 Heterologous expression to enhance enzyme expression and catalytic activity | 在大肠埃希菌中获得了较高的表达量,重组LPMO的表达量达到了600 mg·L-1,在毕赤酵母中则实现了简单快速的分泌表达,利于工业化应用,异源表达后与几丁质酶共同作用于壳聚糖时比色法显示在40 h内产生的还原糖量显著高于野生型组 High expression was achieved in E. coli, with a recombinant LPMO yield of 600 mg·L-1. Simple and rapid secretory expression was realized in P. pastoris, facilitating industrial application. When co-acting with chitinase on chitosan after heterologous expression, the amount of reducing sugar produced within 40 h was significantly higher than that of the wild-type group, as shown by colorimetric method | [ |
| 优化密码子或添加N端标签, 构建含有不同信号肽的表达载体, 提高酶表达量和催化活性 Codon optimization or addition of N-terminal tags, construction of expression vectors with different signal peptides to enhance enzyme expression and catalytic activity | 所有LPMO均在大肠埃希菌周质成功表达,产量至少为40 mg·L-1,在胞质表达的产量在100~800 mg·L-1之间,BatLPMO10产量范围为0.5~2 g·L-1,使用天然信号肽产生的蛋白质量最多,而密码子优化的LsAA9A突变体催化活性高于天然序列,约为天然序列的2倍 All LPMOs were successfully expressed in the periplasm of E. coli, with yields of at least 40 mg·L-1. The yields in cytoplasmic expression ranged from 100 to 800 mg·L-1, and the yield range of BatLPMO10 was 0.5-2 g·L-1. The highest protein production was achieved using the native signal peptide, while the codon-optimized LsAA9A mutant had higher catalytic activity than the native sequence, approximately twice that of the native sequence | [ |
表2 LPMO的基因工程改造结果对比
Table 2 Comparison of LPMO genetic engineering results
| 改造内容 Modification content | 具体结果 Specific result | 参考文献 Reference |
|---|---|---|
| 优化密码子和替换同义密码子, 对LPMO基因进行优化,通过优 化培养条件提高酶表达量,增强 酶活性 Codon optimization and synonymous codon replacement for LPMO gene, enhanced enzyme expression and activity through optimized culture conditions | 优化培养时间、诱导温度与甲醇添加量后,重组毕赤酵母菌株产LPMO能力较优化前增加了12.51%,重组菌株的酶活性最高达到(25.64±0.21) U·L-1;以1%微晶纤维素为底物时,添加LPMO可以使酶解6 h后水解率提升26.46%;以汽爆玉米秸秆为底物时,添加LPMO和一定浓度抗坏血酸可以缩短纤维素酶的降解启动时间,并在酶解9 h后水解效率提升13.07% After optimizing culture time, induction temperature, and methanol addition, the LPMO production of recombinant P. pastoris increased by 12.51% compared with that before optimization. The maximum enzyme activity reached (25.64±0.21) U·L-1. When 1% microcrystalline cellulose was used as the substrate, the hydrolysis rate increased by 26.46% after 6 h of enzymatic hydrolysis with LPMO. When steam-exploded corn stover was used as the substrate, the addition of LPMO and a certain concentration of ascorbic acid shortened the degradation initiation time of cellulase and increased the hydrolysis efficiency by 13.07% after 9 h of enzymatic hydrolysis | [ |
| 构建含有不同信号肽的表达载体, 提高酶表达量 Construction of expression vectors with different signal peptides to enhance enzyme expression | 使用结合了XylS启动子/激活子系统和不同LPMO信号序列的载体,该新型表达系统能可靠产生成熟的LPMO,并且使用SmLPMO10A的信号肽时受试LPMO的表达水平显著提高,SmLPMO10A、BlLPMO10A、JdLPMO10A的产量分别达到22、7、10 mg·L-1 Using a carrier combining the XylS promoter/activator system and different LPMO signal sequences, this novel expression system reliably produces mature LPMO. The expression levels of the tested LPMOs were significantly increased when using the signal peptide of SmLPMO10A, with yields of SmLPMO10A, BlLPMO10A, and JdLPMO10A reaching 22, 7, and 10 mg·L-1, respectively | [ |
| 引入二硫键,提高热稳定性、化学 稳定性和活性 The disulfide bond was introduced to improve the thermal stability, chemical stability and activity | 与野生型相比,突变体在60 ℃下的半衰期增加3倍,表观Tm值上升7 ℃,化学变性中点浓度(midpoint concentration of chemical denaturation, Cm值)提高0.3 mol·L-1,显示出野生型1.5倍的酶活性 Compared with the wild type, the mutant had a 3-fold increased half-life at 60 ℃, a 7 ℃ increase in apparent Tm value, a 0.3 mol·L-1 increase in midpoint concentration of chemical denaturation(Cm value), and 1.5 times the enzyme activity of the wild type | [ |
| 异源表达,提高酶表达量和催化 活性 Heterologous expression to enhance enzyme expression and catalytic activity | 在大肠埃希菌中获得了较高的表达量,重组LPMO的表达量达到了600 mg·L-1,在毕赤酵母中则实现了简单快速的分泌表达,利于工业化应用,异源表达后与几丁质酶共同作用于壳聚糖时比色法显示在40 h内产生的还原糖量显著高于野生型组 High expression was achieved in E. coli, with a recombinant LPMO yield of 600 mg·L-1. Simple and rapid secretory expression was realized in P. pastoris, facilitating industrial application. When co-acting with chitinase on chitosan after heterologous expression, the amount of reducing sugar produced within 40 h was significantly higher than that of the wild-type group, as shown by colorimetric method | [ |
| 优化密码子或添加N端标签, 构建含有不同信号肽的表达载体, 提高酶表达量和催化活性 Codon optimization or addition of N-terminal tags, construction of expression vectors with different signal peptides to enhance enzyme expression and catalytic activity | 所有LPMO均在大肠埃希菌周质成功表达,产量至少为40 mg·L-1,在胞质表达的产量在100~800 mg·L-1之间,BatLPMO10产量范围为0.5~2 g·L-1,使用天然信号肽产生的蛋白质量最多,而密码子优化的LsAA9A突变体催化活性高于天然序列,约为天然序列的2倍 All LPMOs were successfully expressed in the periplasm of E. coli, with yields of at least 40 mg·L-1. The yields in cytoplasmic expression ranged from 100 to 800 mg·L-1, and the yield range of BatLPMO10 was 0.5-2 g·L-1. The highest protein production was achieved using the native signal peptide, while the codon-optimized LsAA9A mutant had higher catalytic activity than the native sequence, approximately twice that of the native sequence | [ |
| [1] | 李欣乐. 秸秆还田对不同土壤团聚体中纤维素降解功能微生物类群的影响[D]. 杨凌: 西北农林科技大学, 2023. |
| LI X L. Effects of straw returning on microbial groups with cellulose degradation function in different soil aggregates[D]. Yangling: Northwest A & F University, 2023. | |
| [2] | WU Z F, LI P F, CHEN Y, et al. Rational design for enhancing cellobiose dehydrogenase activity and its synergistic role in straw degradation[J]. Journal of Agricultural and Food Chemistry, 2024, 72(44): 24620-24631. |
| [3] | 常帅飞, 任文义, 程雨辰, 等. 稻草微贮早期真菌群落演变及其木质纤维素降解酶的预测[J]. 微生物学通报, 2024, 51(5): 1614-1625. |
| CHANG S F, REN W Y, CHENG Y C, et al. Rice straw in early stage of microbial fermentation: fungal community dynamics and lignocellulose-degrading enzymes[J]. Microbiology China, 2024, 51(5): 1614-1625. | |
| [4] | 郑菲, 杨俊钊, 牛羽丰, 等. 嗜热毁丝菌裂解性多糖单加氧酶TtLPMO9I的酶学性质及其功能研究[J]. 生物技术通报, 2024, 40(2): 289-299. |
| ZHENG F, YANG J Z, NIU Y F, et al. Characterization and functional analysis of lytic polysaccharide monooxygenase TtLPMO9I from Thermothelomyces thermophilus[J]. Biotechnology Bulletin, 2024, 40(2): 289-299. | |
| [5] | HEDISON T M, BRESLMAYR E, SHANMUGAM M, et al. Insights into the H2O2-driven catalytic mechanism of fungal lytic polysaccharide monooxygenases[J]. The FEBS Journal, 2021, 288(13): 4115-4128. |
| [6] | BASERA P, CHAKRABORTY S, SHARMA N. Lignocellulosic biomass: insights into enzymatic hydrolysis, influential factors, and economic viability[J]. Discover Sustainability, 2024, 5(1): 311. |
| [7] | VANDHANA T M, REYRE J L, SUSHMAA D, et al. On the expansion of biological functions of lytic polysaccharide monooxygenases[J]. New Phytologist, 2022, 233(6): 2380-2396. |
| [8] | BHATIA S, YADAV S K. Novel catalytic potential of a hyperthermostable mono-copper oxidase (LPMO-AOAA17) for the oxidation of lignin monomers and depolymerisation of lignin dimer in aqueous media[J]. International Journal of Biological Macromolecules, 2021, 186: 563-573. |
| [9] | MENDOZA J A. Novel Aspergillus fumigatus recombinant lpmos: biochemical characteristics, and their participation in saccharification and photobiocatalysis processes[D]. Sao Paulo: University of Sao Paulo, 2022. |
| [10] | ZHANG H Y, ZHOU H C, ZHAO Y, et al. Comparative studies of two AA10 family lytic polysaccharide monooxygenases from Bacillus thuringiensis[J]. PeerJ, 2023, 11: e14670. |
| [11] | SCHRÖDER G C, O’DELL W B, WEBB S P, et al. Capture of activated dioxygen intermediates at the copper-active site of a lytic polysaccharide monooxygenase[J]. Chemical Science, 2022, 13(45): 13303-13320. |
| [12] | HAGEMANN M M, HEDEGÅRD E D. Molecular mechanism of substrate oxidation in lytic polysaccharide monooxygenases: insight from theoretical investigations[J]. Chemistry - A European Journal, 2023, 29(7): e202202379. |
| [13] | BRANDER S, HORVATH I, IPSEN J Ø, et al. Biochemical evidence of both copper chelation and oxygenase activity at the histidine brace[J]. Scientific Reports, 2020, 10(1): 16369. |
| [14] | ZHAO X, XIE F, CHEN K X, et al. The effect of CBM1 and linker on the oxidase, peroxidase and monooxygenase activities of AA9 LPMOs: insight into their correlation with the nature of reductants and crystallinity of celluloses[J]. International Journal of Molecular Sciences, 2024, 25(23): 12616. |
| [15] | HALL K R, MOLLATT M, FORSBERG Z, et al. Impact of the copper second coordination sphere on catalytic performance and substrate specificity of a bacterial lytic polysaccharide monooxygenase[J]. ACS Omega, 2024, 9(21): 23040-23052. |
| [16] | MUNZONE A, EIJSINK V G H, BERRIN J G, et al. Expanding the catalytic landscape of metalloenzymes with lytic polysaccharide monooxygenases[J]. Nature Reviews Chemistry, 2024, 8(2): 106-119. |
| [17] | KUMAR A, SINGH A, SHARMA V K, et al. The upsurge of lytic polysaccharide monooxygenases in biomass deconstruction: characteristic functions and sustainable applications[J]. The FEBS Journal, 2024, 291(23): 5081-5101. |
| [18] | BISSARO B, STREIT B, ISAKSEN I, et al. Molecular mechanism of the chitinolytic peroxygenase reaction[J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(3): 1504-1513. |
| [19] | ØSTBY H, CHRISTENSEN I A, HENNUM K, et al. Functional characterization of a lytic polysaccharide monooxygenase from Schizophyllum commune that degrades non-crystalline substrates[J]. Scientific Reports, 2023, 13: 17373. |
| [20] | WANG B J, ZHANG X, FANG W H, et al. How do metalloproteins tame the Fenton reaction and utilize ·OH radicals in constructive manners[J]. Accounts of Chemical Research, 2022, 55(16): 2280-2290. |
| [21] | KUUSK S, BISSARO B, KUUSK P, et al. Kinetics of H2O2-driven degradation of chitin by a bacterial lytic polysaccharide monooxygenase[J]. Journal of Biological Chemistry, 2018, 293(2): 523-531. |
| [22] | MÜLLER G, CHYLENSKI P, BISSARO B, et al. The impact of hydrogen peroxide supply on LPMO activity and overall saccharification efficiency of a commercial cellulase cocktail[J]. Biotechnology for Biofuels, 2018, 11(1): 209. |
| [23] | YU X R, ZHAO Y, YU J H, et al. Recent advances in the efficient degradation of lignocellulosic metabolic networks by lytic polysaccharide monooxygenase[J]. Acta Biochimica et Biophysica Sinica, 2023, 55(4): 529-539. |
| [24] | IPSEN J Ø, HALLAS-MØLLER M, BRANDER S, et al. Lytic polysaccharide monooxygenases and other histidine-brace copper proteins: structure, oxygen activation and biotechnological applications[J]. Biochemical Society Transactions, 2021, 49(1): 531-540. |
| [25] | TANG C J, GANDLA M L, JÖNSSON L J. LPMO-supported saccharification of biomass: effects of continuous aeration of reaction mixtures with variable fractions of water-insoluble solids and cellulolytic enzymes[J]. Biotechnology for Biofuels and Bioproducts, 2023, 16(1): 156. |
| [26] | FORSBERG Z, BISSARO B, GULLESEN J, et al. Structural determinants of bacterial lytic polysaccharide monooxygenase functionality[J]. Journal of Biological Chemistry, 2018, 293(4): 1397-1412. |
| [27] | 于晓男, 刘霖, 屈明博, 等. 一种几丁质裂解性多糖单加氧酶的活性评价及稳定性研究[J]. 微生物学报, 2022, 62(1): 189-199. |
| YU X N, LIU L, QU M B, et al. The activity and stability analyses of chitin-activity lytic polysaccharide monooxygenase[J]. Acta Microbiologica Sinica, 2022, 62(1): 189-199. | |
| [28] | BORISOVA A S, ISAKSEN T, DIMAROGONA M, et al. Structural and functional characterization of a lytic polysaccharide monooxygenase with broad substrate specificity[J]. Journal of Biological Chemistry, 2015, 290(38): 22955-22969. |
| [29] | LAURENT C V F P, SUN P C, SCHEIBLBRANDNER S, et al. Influence of lytic polysaccharide monooxygenase active site segments on activity and affinity[J]. International Journal of Molecular Sciences, 2019, 20(24): 6219. |
| [30] | CHALAK A, VILLARES A, MOREAU C, et al. Influence of the carbohydrate-binding module on the activity of a fungal AA9 lytic polysaccharide monooxygenase on cellulosic substrates[J]. Biotechnology for Biofuels, 2019, 12(1): 206. |
| [31] | DANNEELS B, TANGHE M, DESMET T. Structural features on the substrate-binding surface of fungal lytic polysaccharide monooxygenases determine their oxidative regioselectivity[J]. Biotechnology Journal, 2019, 14(3): 1800211. |
| [32] | VU V V, BEESON W T, PHILLIPS C M, et al. Determinants of regioselective hydroxylation in the fungal polysaccharide monooxygenases[J]. Journal of the American Chemical Society, 2014, 136(2): 562-565. |
| [33] | LONG L F, HU Y, SUN F B, et al. Advances in lytic polysaccharide monooxygenases with the cellulose-degrading auxiliary activity family 9 to facilitate cellulose degradation for biorefinery[J]. International Journal of Biological Macromolecules, 2022, 219: 68-83. |
| [34] | KOSINAS C, CHOROZIAN K, SANDGREN M, et al. Mutational study of a lytic polysaccharide monooxygenase from Myceliophthora thermophila(MtLPMO9F): structural insights into substrate specificity and regioselectivity[J]. International Journal of Biological Macromolecules, 2025, 288: 138574. |
| [35] | BAGHAEI B, SKRIFVARS M. All-cellulose composites: a review of recent studies on structure, properties and applications[J]. Molecules, 2020, 25(12): 2836. |
| [36] | CHARTI I, SAIR S, RAFIK O, et al. Ecofriendly synthesis of cellulose-silver nanocomposites and the evaluation of their antibacterial activity[J]. Discover Nano, 2025, 20(1): 59. |
| [37] | AYMÉ L, HÉBERT A, HENRISSAT B, et al. Characterization of three bacterial glycoside hydrolase family 9 endoglucanases with different modular architectures isolated from a compost metagenome[J]. Biochimica et Biophysica Acta(BBA) -General Subjects, 2021, 1865(5): 129848. |
| [38] | RAJASEKARAN R, SURESH P K. Physical and chemical methods of extraction of bioactive molecules from Lepidium sativum Linn. and antioxidant activity-based screening and selection of extracts-probable phytochemical, chromatography and mass spectroscopy analysis-based correlates[J]. Research Journal of Pharmacy and Technology, 2021: 3082-3092. |
| [39] | YUAN Y F, WU J, WEI F Q, et al. Elucidating the synergistic action between sulfonated lignin and lytic polysaccharide monooxygenases (LPMOs) in enhancing cellulose hydrolysis[J]. International Journal of Biological Macromolecules, 2025, 296: 139674. |
| [40] | GABER Y, RASHAD B, HUSSEIN R, et al. Heterologous expression of lytic polysaccharide monooxygenases (LPMOs)[J]. Biotechnology Advances, 2020, 43: 107583. |
| [41] | 王逗. Micromonospora aurantiaca几丁质降解酶的挖掘、性质及其协同作用研究[D]. 海口: 海南大学, 2023. |
| WANG D. Study on the identification, properties and synergism of chitinolytic enzymes from Micromonospora aurantiaca[D]. Haikou: Hainan University, 2023. | |
| [42] | NAVARRO D, ROSSO M N, HAON M, et al. Fast solubilization of recalcitrant cellulosic biomass by the basidiomycete fungus Laetisaria arvalis involves successive secretion of oxidative and hydrolytic enzymes[J]. Biotechnology for Biofuels, 2014, 7(1): 143. |
| [43] | 李桐冰. 红侧耳裂解多糖单加氧酶基因PdLPMO9A的克隆表达及其酶学性质表征[D]. 长春: 吉林农业大学, 2021. |
| LI T B. Cloning, expression and characterization of lytic polysaccharide monooxygenase gene PdLPMO9A from Pleurotus djamor[D]. Changchun: Jilin Agricultural University, 2021. | |
| [44] | LI Y L, LI T B, GUO J T, et al. Expression and characterization of a novel lytic polysaccharide monooxygenase, PdLPMO9A, from the edible fungus Pleurotus djamor and its synergistic interactions with cellulase in corn straw biomass saccharification[J]. Bioresource Technology, 2022, 348: 126792. |
| [45] | AGRAWAL R, VERMA A, SINGHANIA R R, et al. Current understanding of the inhibition factors and their mechanism of action for the lignocellulosic biomass hydrolysis[J]. Bioresource Technology, 2021, 332: 125042. |
| [46] | FORSBERG Z, STEPNOV A A, NÆRDAL G K, et al. Engineering lytic polysaccharide monooxygenases (LPMOs)[J]. Methods in Enzymology, 2020, 644: 1-34. |
| [47] | CZJZEK M, FICKO-BLEAN E, BERRIN J G. A special issue of Essays in Biochemistry on current advances about CAZymes and their impact and key role in human health and environment[J]. Essays in Biochemistry, 2023, 67(3): 325-329. |
| [48] | EIBINGER M, GANNER T, BUBNER P, et al. Cellulose surface degradation by a lytic polysaccharide monooxygenase and its effect on cellulase hydrolytic efficiency[J]. Journal of Biological Chemistry, 2014, 289(52): 35929-35938. |
| [49] | 梁丛颖, 林璐. 环境微生物介导的木质素代谢及其资源化利用研究进展[J]. 微生物学通报, 2020, 47(10): 3380-3392. |
| LIANG C Y, LIN L. Environmental microorganisms driven lignin biodegradation and their roles in lignin utilization[J]. Microbiology China, 2020, 47(10): 3380-3392. | |
| [50] | LI F, ZHANG J L, MA F Y, et al. Lytic polysaccharide monooxygenases promote oxidative cleavage of lignin and lignin-carbohydrate complexes during fungal degradation of lignocellulose[J]. Environmental Microbiology, 2021, 23(8): 4547-4560. |
| [51] | SUN S, LI F, LI M Z, et al. Lytic polysaccharide monooxygenase synergized with lignin-degrading enzymes for efficient lignin degradation[J]. iScience, 2023, 26(10): 107870. |
| [52] | CHEN K X, ZHAO X, ZHANG P Y, et al. A novel AA14 LPMO from Talaromyces rugulosus with bifunctional cellulolytic/hemicellulolytic activity boosted cellulose hydrolysis[J]. Biotechnology for Biofuels and Bioproducts, 2024, 17(1): 30. |
| [53] | ZHANG M J, FU X P, GU R R, et al. A novel starch-active lytic polysaccharide monooxygenase discovered with bioinformatics screening and its application in textile desizing[J]. BMC Biotechnology, 2024, 24(1): 2. |
| [54] | ESKANDARI A, NEZHAD N G, LEOW T C, et al. Current achievements, strategies, obstacles, and overcoming the challenges of the protein engineering in Pichia pastoris expression system[J]. World Journal of Microbiology and Biotechnology, 2023, 40(1): 39. |
| [55] | BATTH T S, SIMONSEN J L, HERNÁNDEZ-ROLLÁN C, et al. A seven-transmembrane methyltransferase catalysing N-terminal histidine methylation of lytic polysaccharide monooxygenases[J]. Nature Communications, 2023, 14: 4202. |
| [56] | 马立娟, 佟文哲, 杜丽平, 等. N-糖基化对黑曲霉AnLPMO15g与纤维素酶协同作用的影响[J]. 高校化学工程学报, 2019, 33(5): 1148-1155. |
| MA L J, TONG W Z, DU L P, et al. Effects of N-glycosylation on synergism of AnLPMO15g from Aspergillus niger and cellulase[J]. Journal of Chemical Engineering of Chinese Universities, 2019, 33(5): 1148-1155. | |
| [57] | 王旭昕. 裂解多糖单加氧酶在巴斯德毕赤酵母的异源表达及其应用[D]. 呼和浩特: 内蒙古大学, 2023. |
| WANG X X. Heterologous expression of lytic polysaccharide monooxygenase in Pichia pastoris and its applications[D]. Hohhot: Inner Mongolia University, 2023. | |
| [58] | KADOWAKI M A S, VÁRNAI A, JAMESON J K, et al. Functional characterization of a lytic polysaccharide monooxygenase from the thermophilic fungus Myceliophthora thermophila[J]. PLoS One, 2018, 13(8): e0202148. |
| [59] | COURTADE G, LE S B, SÆTROM G I, et al. A novel expression system for lytic polysaccharide monooxygenases[J]. Carbohydrate Research, 2017, 448: 212-219. |
| [60] | RUSSO D A, ZEDLER J A Z, WITTMANN D N, et al. Expression and secretion of a lytic polysaccharide monooxygenase by a fast-growing Cyanobacterium[J]. Biotechnology for Biofuels, 2019, 12(1): 74. |
| [61] | ZHOU X L, XU Z Q, LI Y Q, et al. Improvement of the stability and activity of an LPMO through rational disulfide bonds design[J]. Frontiers in Bioengineering and Biotechnology, 2022, 9: 815990. |
| [62] | 朱铃. Aspergillus oryzae溶解多糖单加氧酶基因的克隆,异源表达及酶学特性分析[D]. 昆明: 云南大学, 2016. |
| ZHU L. Gene cloning, heteroexpression and characterization of the lytic polysaccharide monooxygenase from Aspergillus oryzae[D]. Kunming: Yunnan University, 2016. | |
| [63] | HERNÁNDEZ-ROLLÁN C, FALKENBERG K B, RENNIG M, et al. LyGo: a platform for rapid screening of lytic polysaccharide monooxygenase production[J]. ACS Synthetic Biology, 2021, 10(4): 897-906. |
| [64] | 黄天晨, 杨建花, 唐梓静, 等. 裂解性多糖单加氧酶HcLPMO与纤维素酶协同降解纤维素[J]. 微生物学报, 2021, 61(7): 1971-1982. |
| HUANG T C, YANG J H, TANG Z J, et al. Synergistic degradation of cellulose by lytic polysaccharides monooxygenase HcLPMO and cellulase[J]. Acta Microbiologica Sinica, 2021, 61(7): 1971-1982. | |
| [65] | HSU J M, LI C W, LAI Y J, et al. Posttranslational modifications of PD-L1 and their applications in cancer therapy[J]. Cancer Research, 2018, 78(22): 6349-6353. |
| [66] | GUO X, AN Y J, CHAI C C, et al. Construction of the R17L mutant of MtC1LPMO for improved lignocellulosic biomass conversion by rational point mutation and investigation of the mechanism by molecular dynamics simulations[J]. Bioresource Technology, 2020, 317: 124024. |
| [67] | CHENG C, HAIDER J, LIU P, et al. Engineered LPMO significantly boosting cellulase-catalyzed depolymerization of cellulose[J]. Journal of Agricultural and Food Chemistry, 2020, 68(51): 15257-15266. |
| [68] | JENSEN M S, KLINKENBERG G, BISSARO B, et al. Engineering chitinolytic activity into a cellulose-active lytic polysaccharide monooxygenase provides insights into substrate specificity[J]. Journal of Biological Chemistry, 2019, 294(50): 19349-19364. |
| [69] | ZARATTINI M, CORSO M, KADOWAKI M A, et al. LPMO-oxidized cellulose oligosaccharides evoke immunity in Arabidopsis conferring resistance towards necrotrophic fungus B. cinerea[J]. Communications Biology, 2021, 4: 727. |
| [70] | 周頔, 徐登薇, 宋锐, 等. 低聚糖类益生元对炼乳品质的影响[J]. 食品安全质量检测学报, 2022, 13(18): 5887-5894. |
| ZHOU D, XU D W, SONG R, et al. Effects of oligosaccharide prebiotics on the quality of condensed milk[J]. Journal of Food Safety & Quality, 2022, 13(18): 5887-5894. | |
| [71] | WENINGER S N, HERMAN C, MEYER R K, et al. Oligofructose improves small intestinal lipid-sensing mechanisms via alterations to the small intestinal microbiota[J]. Microbiome, 2023, 11(1): 169. |
| [72] | YANG C C, DU Y, LI Q M, et al. Fructo-oligosaccharides alleviated ulcerative colitis via gut microbiota-dependent tryptophan metabolism in association with aromatic hydrocarbon receptor activation in mice[J]. Journal of Agricultural and Food Chemistry, 2024, 72(50): 27912-27922. |
| [73] | 刘重慧, 张静, 范誉川, 等. 低聚异麦芽糖调节肠道菌群及润肠通便作用的研究[J]. 食品工业科技, 2018, 39(7): 298-302. |
| LIU C H, ZHANG J, FAN Y C, et al. Studies on the effects of isomaltooligosaccharides regulating intestinal microbial flora proliferation and relieving constipated function[J]. Science and Technology of Food Industry, 2018, 39(7): 298-302. | |
| [74] | 周火祥, 李倩, 王满, 等. 基于调节肠道菌群的功能性低聚糖改善肥胖的研究进展[J]. 中国微生态学杂志, 2022, 34(11): 1360-1364. |
| ZHOU H X, LI Q, WANG M, et al. Amelioration of obesity by functional oligosaccharides based on regulatory role of gut microbiota: research progress[J]. Chinese Journal of Microecology, 2022, 34(11): 1360-1364. | |
| [75] | WANG W B, NIE Y S, TIAN H, et al. Microbial community, co-occurrence network relationship and fermentation lignocellulose characteristics of Broussonetia papyrifera ensiled with wheat bran[J]. Microorganisms, 2022, 10(10): 2015. |
| [76] | CHEN Y X, SHEN D, ZHANG L L, et al. Supplementation of non-starch polysaccharide enzymes cocktail in a corn-miscellaneous meal diet improves nutrient digestibility and reduces carbon dioxide emissions in finishing pigs[J]. Animals, 2020, 10(2): 232. |
| [1] | 韦庆翠, 姜娜英, 沈骏扬, 张焕朝, 张衡锋. 化肥减量配施生物质炭对高沙土氮磷淋失及土壤性质的影响[J]. 浙江农业学报, 2025, 37(9): 1943-1950. |
| [2] | 江涛, 王立国, 孙芳芳, 成剑波, 何腾兵, 秦松, 范成五, 阴文芳. 沼渣生物质炭对西南喀斯特山区沼液灌溉土壤氮淋溶和白菜产量的影响[J]. 浙江农业学报, 2021, 33(11): 2104-2115. |
| [3] | 李倩, 许之扬, 阮文权. 黄孢原毛平革菌后处理深度提升醋糟产甲烷潜力[J]. 浙江农业学报, 2020, 32(5): 904-911. |
| [4] | 陈重军, 刘玉学, 冯宇, 王建芳. 添加竹炭对设施菜地土壤氮磷流失和微生物群落结构的影响[J]. 浙江农业学报, 2018, 30(1): 123-128. |
| [5] | 陈红卫, 黄玲, 冯露, 李晓庆, 孟雨田, 代琳. 生物质炭基肥对农田土壤温室气体排放年际变化的影响[J]. 浙江农业学报, 2017, 29(6): 977-981. |
| [6] | 林肖庆1,2,吕豪豪2,3,刘玉学2,3,汪玉瑛2,3,杨生茂1,2,3,*. 生物质原料及炭化温度对生物炭产率与性质的影响[J]. 浙江农业学报, 2016, 28(7): 1216-. |
| [7] | 代琳, 聂颖, 冯露, 许毛毛, 丁弈君, 徐红涛, 王宏燕. 生物质炭施入对白浆土碳氮变化的影响[J]. 浙江农业学报, 2016, 28(10): 1745-1754. |
| [8] | 田伟;张振华;汪贞;王霞;李妍;刘振华*. 牛粪高温堆肥过程中木质纤维素降解及相关生物学特性研究[J]. , 2014, 26(2): 0-432438. |
| [9] | 陈江;章旭明. 村镇垃圾混合生物质衍生燃料(RDF)燃烧性能及氯化氢排放特性研究[J]. , 2011, 23(3): 0-581. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||

