浙江农业学报 ›› 2026, Vol. 38 ›› Issue (2): 284-300.DOI: 10.3969/j.issn.1004-1524.20250193
青钱柳叶片叶绿体基因组结构特征与系统发育分析
尹明华1,2( ), 康海燕1, 钟可情1, 吴利鹃3,4, 胡烈新3,4, 吴春发3,4, 龚水飞3,4
), 康海燕1, 钟可情1, 吴利鹃3,4, 胡烈新3,4, 吴春发3,4, 龚水飞3,4
1.上饶师范学院 生命科学学院 江西 上饶 334001 2.上饶市药食同源植物资源保护与利用重点实验室 江西 上饶 334001 3.江西琪格轩生态农业开发有限公司 江西 上饶 334699 4.上饶县琪轩种植专业合作社 江西 上饶 334699
-
收稿日期:2025-03-12出版日期:2026-02-25发布日期:2026-03-24 -
作者简介:尹明华,研究方向为药用植物生物技术。E-mail:yinminghua04@163.com -
基金资助:国家自然科学基金(31960079);国家自然科学基金(32060092);江西省现代农业产业技术体系建设专项(JXARS-13-赣东站)
Structural characterization and phylogenetic analysis of the chloroplast genome of Cyclocarya paliurus (Batal.) Iljinskaja leaves
YIN Minghua1,2( ), KANG Haiyan1, ZHONG Keqing1, WU Lijuan3,4, HU Liexin3,4, WU Chunfa3,4, GONG Shuifei3,4
), KANG Haiyan1, ZHONG Keqing1, WU Lijuan3,4, HU Liexin3,4, WU Chunfa3,4, GONG Shuifei3,4
1. College of Life Sciences ,Shangrao Normal University Shangrao 334001, Jiangxi, China 2. Shangrao Key Laboratory of Protection and Utilization of Medicinal and Edible Plant Resources Shangrao 334001, Jiangxi, China 3. Jiangxi Qigexuan Ecological Agriculture Development Co. ,Ltd. Shangrao 334699, Jiangxi, China 4. Shangrao Qixuan Planting Professional Cooperative Shangrao 334699, Jiangxi, China
-
Received:2025-03-12Published:2026-02-25Online:2026-03-24
摘要:
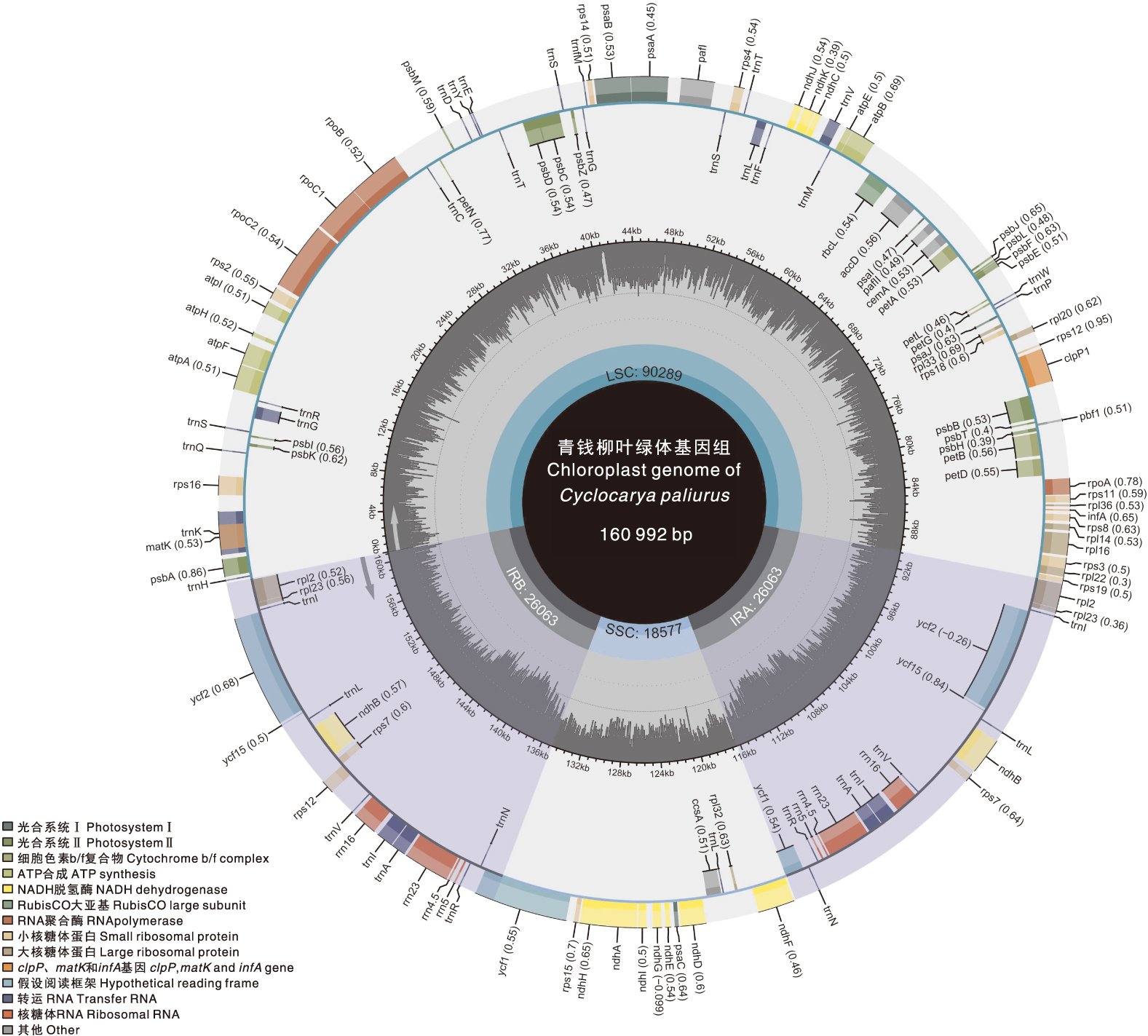
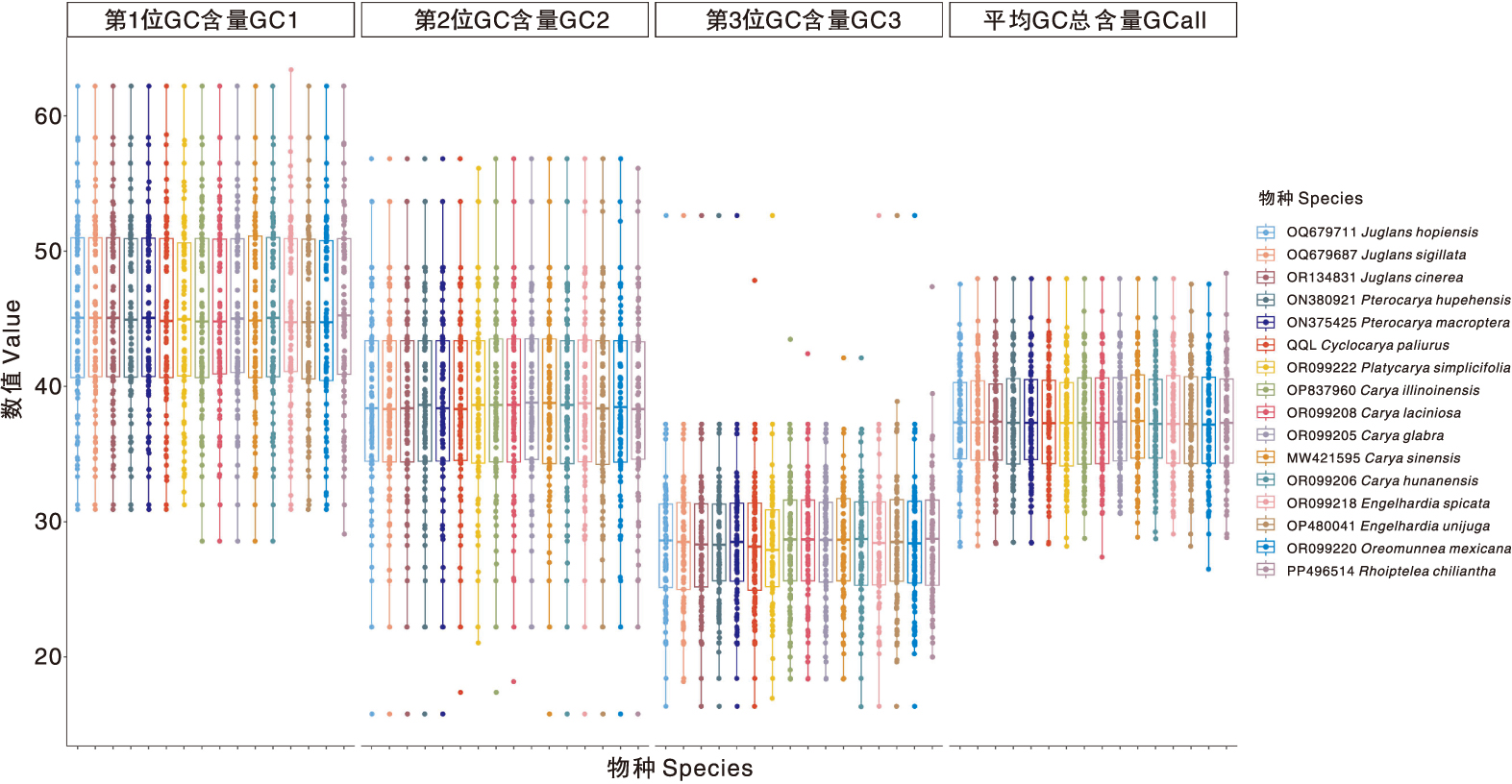
为探究青钱柳[Cyclocarya paliurus(Batal.) Iljinskaja]叶绿体基因组的结构特征与系统进化关系,本研究以江西省上饶市广信区青钱柳叶片为材料,通过高通量测序获得其叶绿体基因组序列,利用生物信息学方法分析其序列特征、密码子使用偏好性与系统发育关系。结果显示,上饶市广信区青钱柳叶绿体基因组全长为160 992 bp,平均GC含量为37.48%;共注释到133个基因,包括88个蛋白质编码基因、37个tRNA基因和8个rRNA基因;鉴定到84个简单重复序列(SSR),以A/T重复为主;52个长重复序列,以正向重复和回文重复为主。与15个近缘种比较显示,青钱柳叶绿体基因组在LSC-IRB边界(JLB)、SSC-IRB边界(JSB)、SSC-IRA边界(JSA)和LSC-IRA边界(JLA)4个边界区域的基因类型与排列顺序基本一致,边界扩张与收缩程度差异不明显;在LSC和SSC共鉴定出10个高变异区域,分别为matK_rps16、rps16_trnQ-UUG、psbK_psbI、trnS-GCU_trnG-GCC、trnT-GGU_psbD、ndhC_trnV-UAC、trnV-UAC、rpl22、ycf1和rps15_ycf1。青钱柳叶绿体基因组密码子整体偏好性较弱,主要受自然选择影响;共鉴定出15个最优密码子:AAU、GAA、CAU、AAA、AUU、AUG、GGA、GGU、CCU、ACA、GUA、GUU、CGU、UUG和AGU,多数以U或A结尾。系统发育分析表明,上饶市广信区青钱柳与南京青钱柳二倍体(MW531677,Cyclocarya paliurus isolate 2nPA)聚为一支,亲缘关系较近。本研究结果可为青钱柳遗传多样性研究、分子育种和物种鉴定提供理论依据。
中图分类号:
引用本文
尹明华, 康海燕, 钟可情, 吴利鹃, 胡烈新, 吴春发, 龚水飞. 青钱柳叶片叶绿体基因组结构特征与系统发育分析[J]. 浙江农业学报, 2026, 38(2): 284-300.
YIN Minghua, KANG Haiyan, ZHONG Keqing, WU Lijuan, HU Liexin, WU Chunfa, GONG Shuifei. Structural characterization and phylogenetic analysis of the chloroplast genome of Cyclocarya paliurus (Batal.) Iljinskaja leaves[J]. Acta Agriculturae Zhejiangensis, 2026, 38(2): 284-300.
| 基因功能 Gene function | 基因类型 Gene type | 基因名 Gene name | 基因数量 Number of genes |
|---|---|---|---|
| 光合作用 | 光系统Ⅰ Photosystem Ⅰ | psaA、psaB、psaC、psaI、psaJ | 5 |
| Photosynthesis | 光系统Ⅱ Photosystem Ⅱ | psbA、psbB、psbC、psbD、psbE、psbF、psbH、psbI、psaJ、psbK、psbL、psbM、psbT、psbZ、psbN(pbf1) | 15 |
| NADH脱氢酶 NADH dehydrogenase | ndhA、ndhB*、ndhC、ndhD、ndhE、ndhF、ndhG、ndhH、ndhI、ndhJ、ndhK | 12 | |
| 细胞色素b/f复合体 Cytochrome b/f complex | petA、petB、petD、petG、petL、petN | 6 | |
| ATP合成酶ATP synthase | atpA、atpB、atpE、atpF、atpH、atpI | 6 | |
| 自我复制 Self replication | 核糖体大亚基蛋白质 Ribosomal large subunit protein | rpl14、rpl16、rpl2*、rpl20、rpl22、rpl23*、rpl32、rpl33、rpl36 | 11 |
| 核糖体小亚基蛋白质 Ribosomal small subunit protein | rps11、rps12*、rps14、rps15、rps16、rps18、rps19、rps2、rps3、rps4、rps7*、rps8 | 14 | |
| 核糖体大亚基Ribosomal large subunit | rbcL | 1 | |
| RNA聚合酶RNA polymerase | rpoA、rpoB、rpoC1、rpoC2 | 4 | |
| 核糖体RNA Ribosomal RNA | rrn16*、rrn23*、rrn4.5*、rrn5* | 8 | |
| 转运RNA transfer RNA | trnA-UGC*、trnC-GCA、trnD-GUC、trnE-UUC、trnF-GAA、trnfM-CAU、trnG-GCC、trnG-UCC、trnH-GUG、trnI-CAU*、trnI-GAU*、trnK-UUU、trnL-CAA*、trnL-UAA、trnL-UAG、trnM-CAU、trnN-GUU*、trnP-UGG、trnQ-UUG、trnR-ACG*、trnR-UCU、trnS-GCU、trnS-GGA、trnS-UGA、trnT-GGU、trnT-UGU、trnV-GAC*、trnV-UAC、trnW-CCA、trnY-GUA | 37 | |
| 其他基因Other genes | 成熟酶Mature enzyme | matK | 1 |
| 蛋白酶Protease | clpP1 | 1 | |
| 囊膜蛋白Envelope protein | cemA | 1 | |
| 乙酰辅酶A羧化酶 Acetyl-CoA carboxylase | accD | 1 | |
| C-型细胞色素合成基因 C-type cytochrome synthesis gene | ccsA | 1 | |
| 翻译起始因子Translation initiation factor | infA | 1 | |
| 未知功能基因 Unknown genes | 保守假设叶绿体阅读框架 Conservative assumption of chloroplast reading framework | ycf1*、ycf15*、ycf2*、ycf3(pafI)、ycf4(pafII) | 8 |
表1 青钱柳叶绿体基因组编码基因
Table 1 Genes encoded by the chloroplast genome of Cyclocarya paliurus(Batal.) Iljinsk
| 基因功能 Gene function | 基因类型 Gene type | 基因名 Gene name | 基因数量 Number of genes |
|---|---|---|---|
| 光合作用 | 光系统Ⅰ Photosystem Ⅰ | psaA、psaB、psaC、psaI、psaJ | 5 |
| Photosynthesis | 光系统Ⅱ Photosystem Ⅱ | psbA、psbB、psbC、psbD、psbE、psbF、psbH、psbI、psaJ、psbK、psbL、psbM、psbT、psbZ、psbN(pbf1) | 15 |
| NADH脱氢酶 NADH dehydrogenase | ndhA、ndhB*、ndhC、ndhD、ndhE、ndhF、ndhG、ndhH、ndhI、ndhJ、ndhK | 12 | |
| 细胞色素b/f复合体 Cytochrome b/f complex | petA、petB、petD、petG、petL、petN | 6 | |
| ATP合成酶ATP synthase | atpA、atpB、atpE、atpF、atpH、atpI | 6 | |
| 自我复制 Self replication | 核糖体大亚基蛋白质 Ribosomal large subunit protein | rpl14、rpl16、rpl2*、rpl20、rpl22、rpl23*、rpl32、rpl33、rpl36 | 11 |
| 核糖体小亚基蛋白质 Ribosomal small subunit protein | rps11、rps12*、rps14、rps15、rps16、rps18、rps19、rps2、rps3、rps4、rps7*、rps8 | 14 | |
| 核糖体大亚基Ribosomal large subunit | rbcL | 1 | |
| RNA聚合酶RNA polymerase | rpoA、rpoB、rpoC1、rpoC2 | 4 | |
| 核糖体RNA Ribosomal RNA | rrn16*、rrn23*、rrn4.5*、rrn5* | 8 | |
| 转运RNA transfer RNA | trnA-UGC*、trnC-GCA、trnD-GUC、trnE-UUC、trnF-GAA、trnfM-CAU、trnG-GCC、trnG-UCC、trnH-GUG、trnI-CAU*、trnI-GAU*、trnK-UUU、trnL-CAA*、trnL-UAA、trnL-UAG、trnM-CAU、trnN-GUU*、trnP-UGG、trnQ-UUG、trnR-ACG*、trnR-UCU、trnS-GCU、trnS-GGA、trnS-UGA、trnT-GGU、trnT-UGU、trnV-GAC*、trnV-UAC、trnW-CCA、trnY-GUA | 37 | |
| 其他基因Other genes | 成熟酶Mature enzyme | matK | 1 |
| 蛋白酶Protease | clpP1 | 1 | |
| 囊膜蛋白Envelope protein | cemA | 1 | |
| 乙酰辅酶A羧化酶 Acetyl-CoA carboxylase | accD | 1 | |
| C-型细胞色素合成基因 C-type cytochrome synthesis gene | ccsA | 1 | |
| 翻译起始因子Translation initiation factor | infA | 1 | |
| 未知功能基因 Unknown genes | 保守假设叶绿体阅读框架 Conservative assumption of chloroplast reading framework | ycf1*、ycf15*、ycf2*、ycf3(pafI)、ycf4(pafII) | 8 |
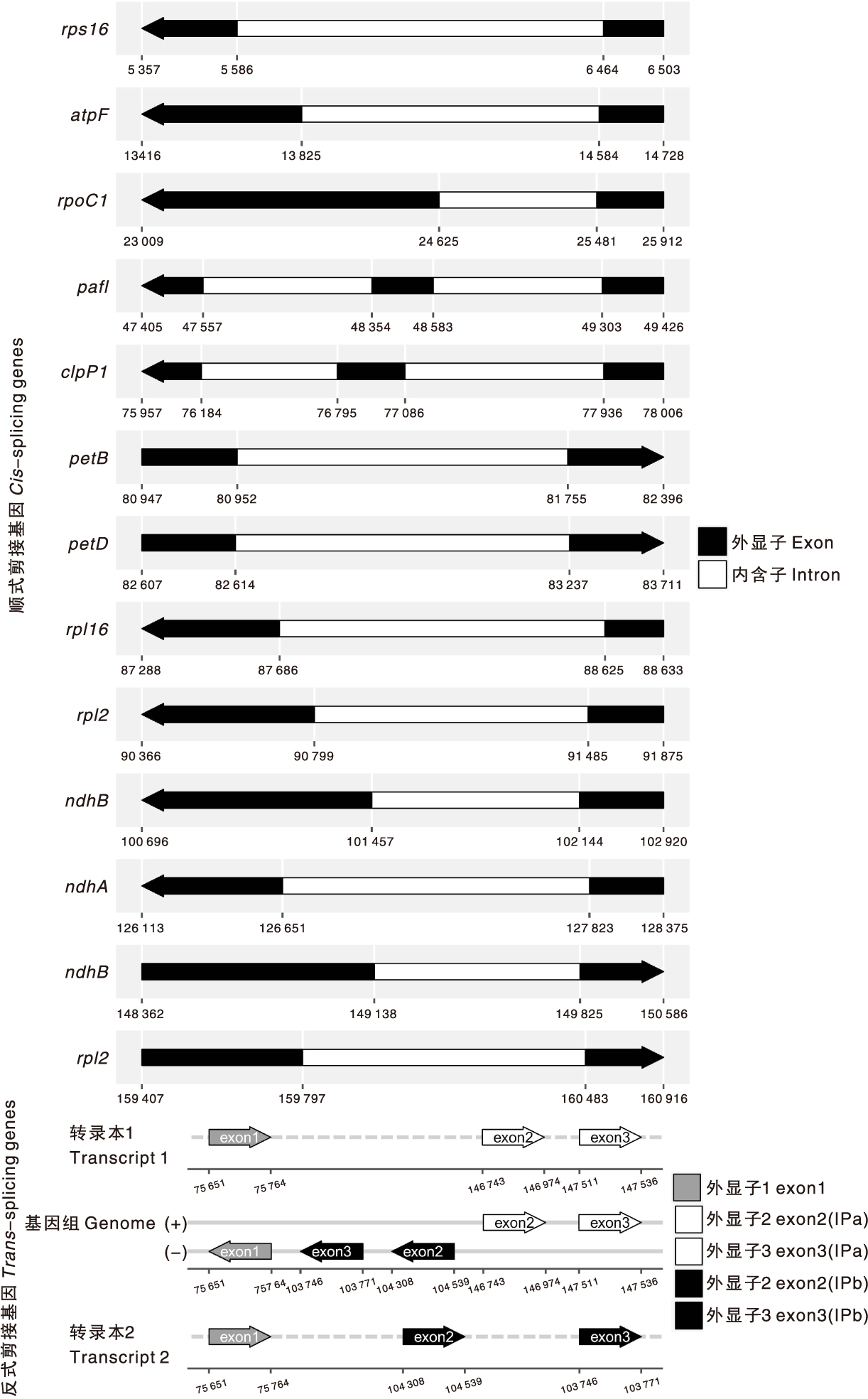

图2 青钱柳叶绿体基因组顺式剪接基因与反式剪接基因示意图
Fig.2 The cis-splicing gene map and the trans-splicing gene map of chloroplast genome of Cyclocarya paliurus(Batal.) Iljinsk
| 重复单元碱基类型 Repetitive unit base type | 重复单元重复次数Repetitive unit repetition times | 总数 Total number | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | ||
| A/T | 0 | 0 | 0 | 0 | 0 | 27 | 20 | 13 | 5 | 6 | 0 | 1 | 1 | 73 |
| C/G | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 |
| AT/AT | 0 | 2 | 4 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 8 |
| AAT/ATT | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
表2 青钱柳叶绿体基因组SSR分布
Table 2 Distribution of SSRs in the chloroplast genome of Cyclocarya paliurus(Batal.) Iljinsk
| 重复单元碱基类型 Repetitive unit base type | 重复单元重复次数Repetitive unit repetition times | 总数 Total number | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | ||
| A/T | 0 | 0 | 0 | 0 | 0 | 27 | 20 | 13 | 5 | 6 | 0 | 1 | 1 | 73 |
| C/G | 0 | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 |
| AT/AT | 0 | 2 | 4 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 8 |
| AAT/ATT | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
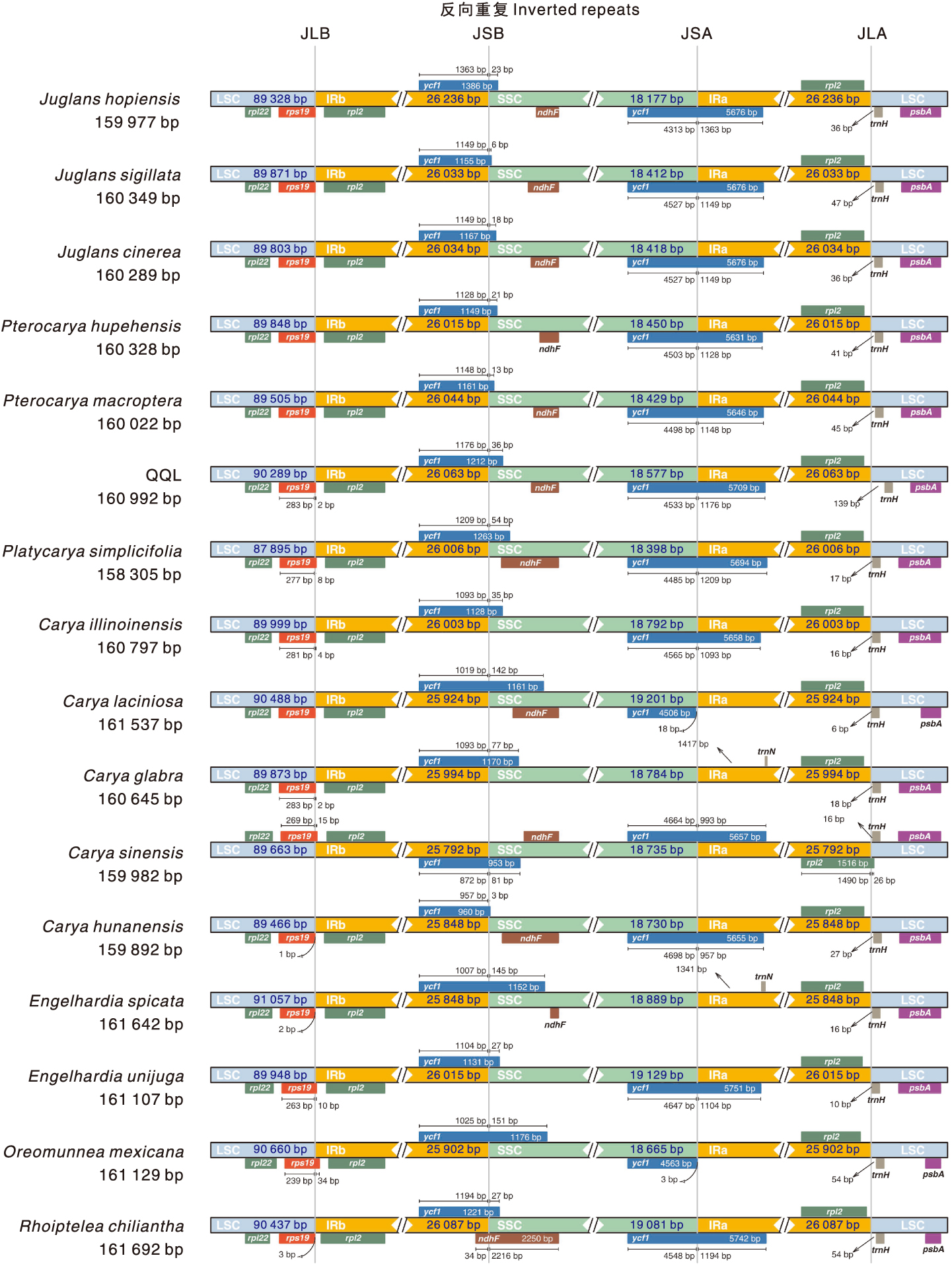
图3 青钱柳与15个近缘种叶绿体基因组IR边界的结构差异
Fig.3 Structural differences of the inverted repeat (IR) boundaries in the chloroplast genomes of Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species
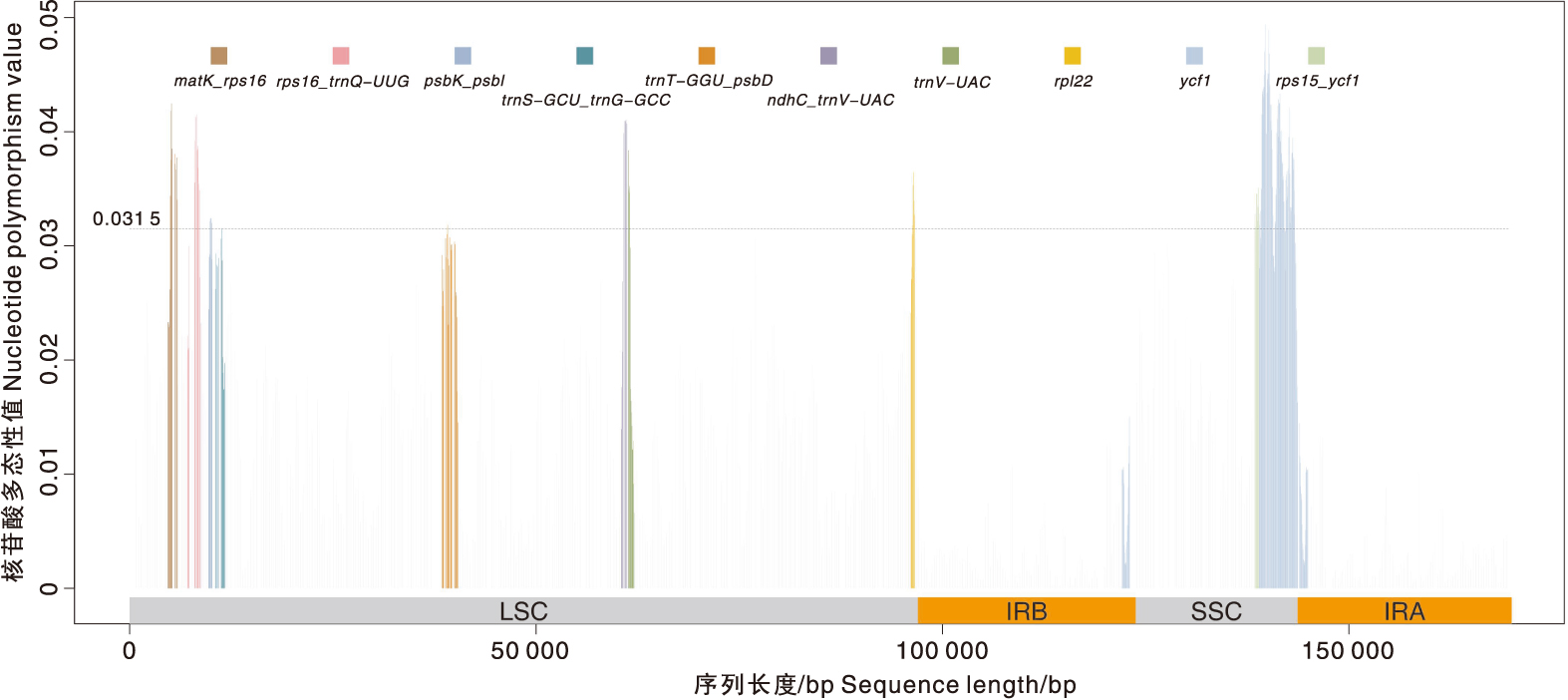
图4 青钱柳和15个近缘种叶绿体基因组的核苷酸多态性
Fig.4 Nucleotide polymorphism (Pi) in chloroplast genomes of Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species
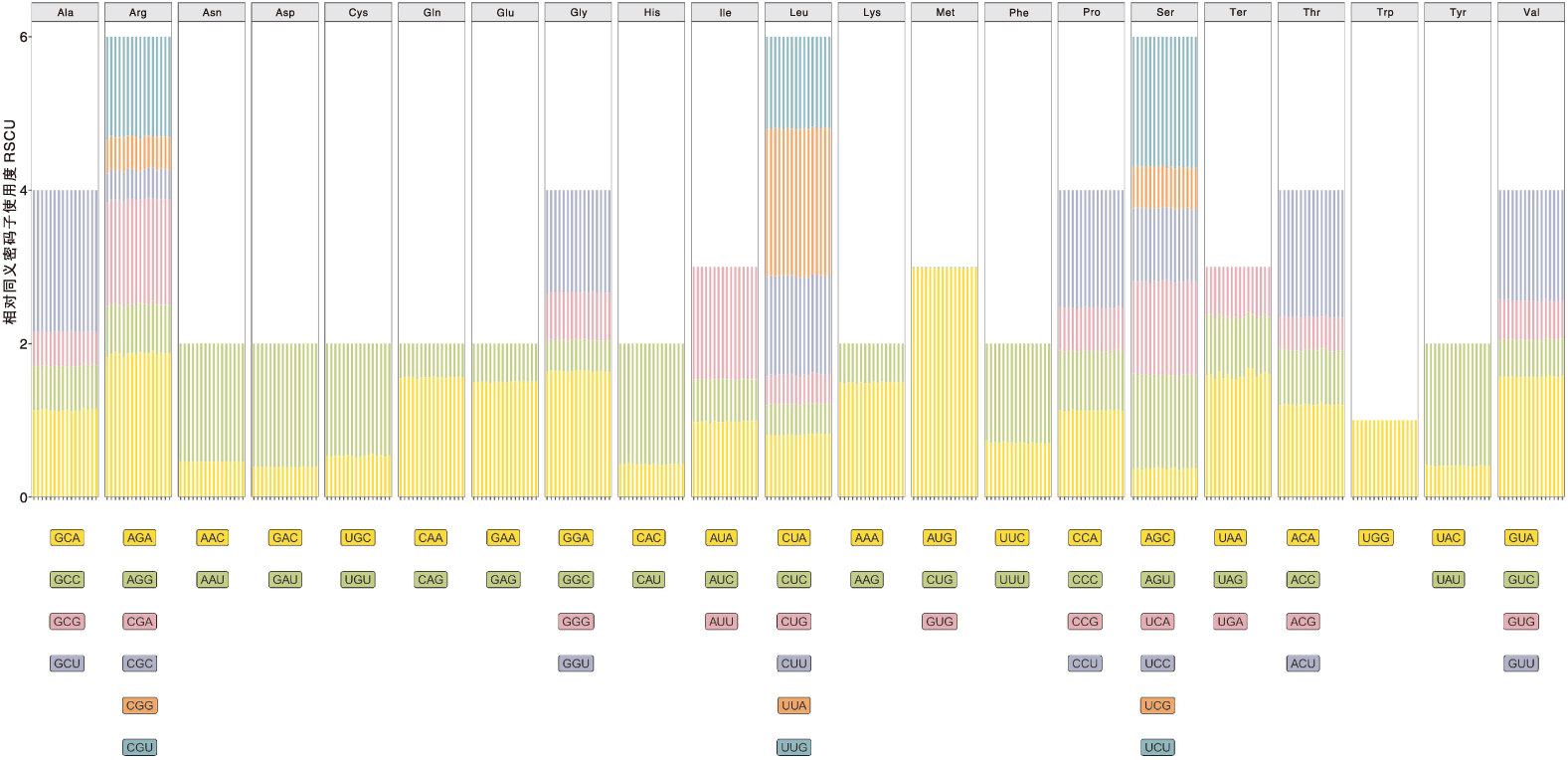
图6 青钱柳与15个近缘种叶绿体基因组的相对同义密码子使用度 图中每个氨基酸从左到右的竖线表示青钱柳和15个近缘种,16个品种的排序与图5中的排序一致。
Fig.6 Relative synonymous codon usage of chloroplast genome codons in Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species The vertical lines from left to right in each amino acid in the figure represent Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species. The ranking of the 16 varieties is consistent with the ranking of the 16 varieties in Fig. 5.
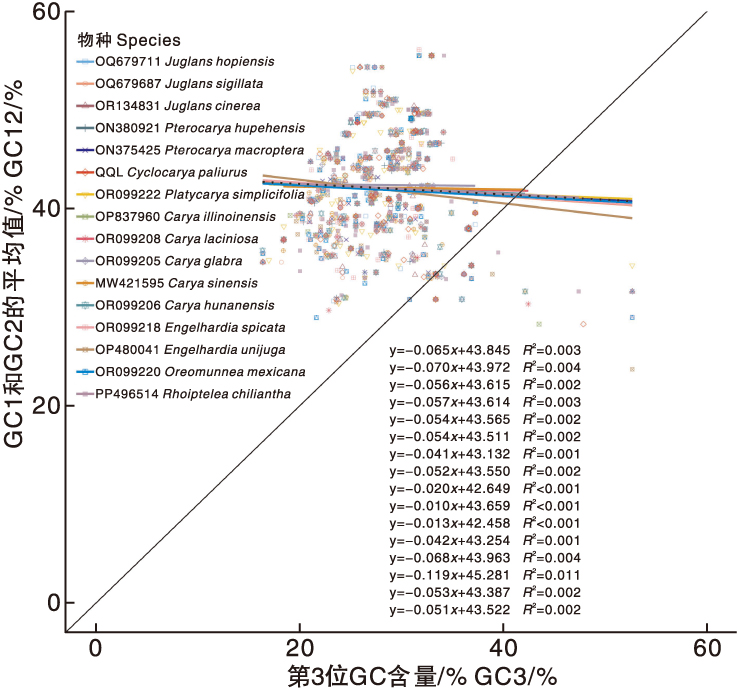
图7 青钱柳与15个近缘种叶绿体基因组密码子的中性绘图分析
Fig.7 Neutrality plot analysis of chloroplast genome codons in Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species
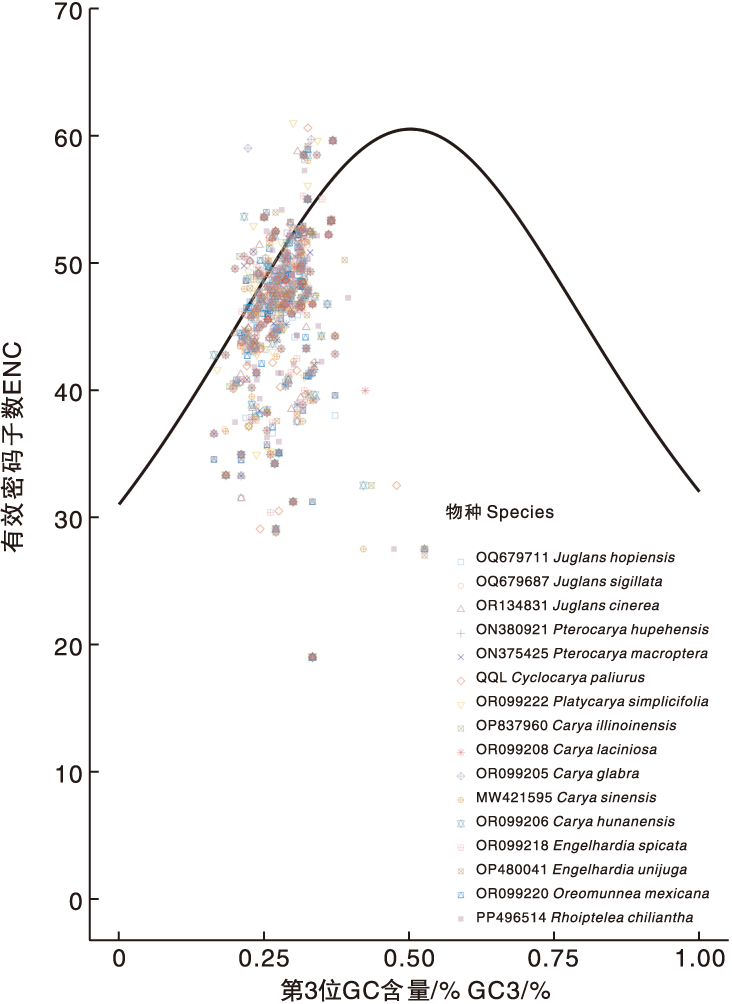
图8 青钱柳与15个近缘种叶绿体基因组密码子使用偏好性的ENC-plot分析
Fig.8 ENC-plot analysis of chloroplast genome codons in Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species
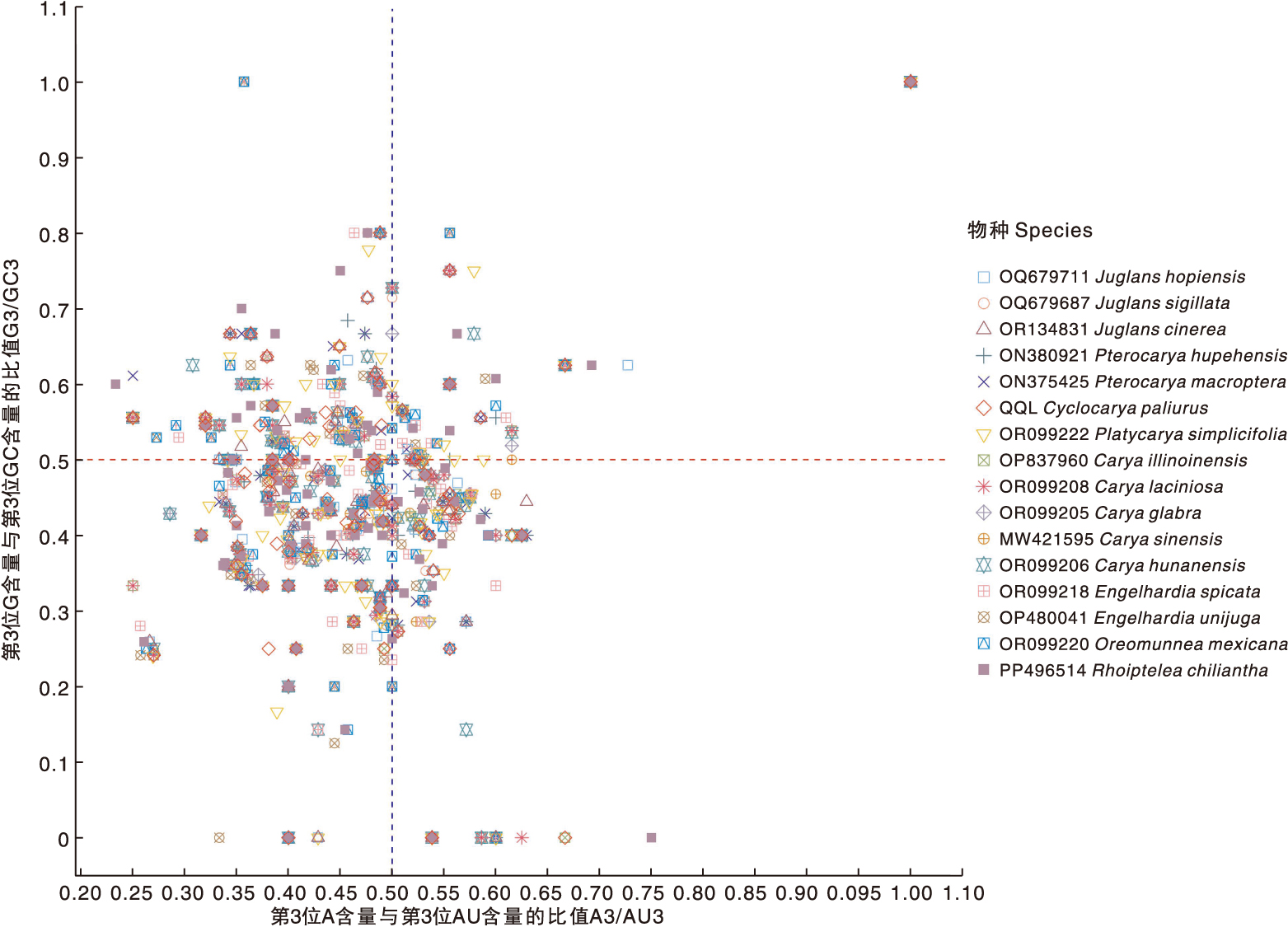
图9 青钱柳与15个近缘种叶绿体基因组密码子的PR2-plot分析
Fig.9 PR2-plot analysis of chloroplast genomes codons in Cyclocarya paliurus(Batal.) Iljinsk and 15 closely related species
| 密码子 Codon | 氨基酸 Amino acid | 相对同义密码子使用度 Relative synonymous codon usage | RSCU低表达 RSCU low expression | RSCU高表达 RSCU high expression | RSCU差值 RSCU difference |
|---|---|---|---|---|---|
| AAU | 天冬酰胺Asn | 1.535 | 1.375 | 1.500 | 0.125 |
| GAA | 谷氨酸Glu | 1.508 | 1.556 | 1.714 | 0.159 |
| CAU | 组氨酸His | 1.574 | 1.000 | 1.200 | 0.200 |
| AAA | 赖氨酸Lys | 1.494 | 1.000 | 1.333 | 0.333 |
| AUU | 异亮氨酸Ile | 1.461 | 1.200 | 1.800 | 0.600 |
| AUG | 蛋氨酸Met | 2.981 | 1.000 | 1.556 | 0.556 |
| GGA | 甘氨酸Gly | 1.634 | 1.647 | 1.846 | 0.199 |
| GGU | 甘氨酸Gly | 1.333 | 0.706 | 0.923 | 0.217 |
| CCU | 脯氨酸Pro | 1.502 | 1.143 | 1.500 | 0.357 |
| ACA | 苏氨酸Thr | 1.213 | 1.091 | 1.500 | 0.409 |
| GUA | 缬氨酸Val | 1.566 | 0.923 | 2.000 | 1.077 |
| GUU | 缬氨酸Val | 1.445 | 1.154 | 1.333 | 0.179 |
| CGU | 精氨酸Arg | 1.304 | 0.278 | 0.545 | 0.268 |
| UUG | 亮氨酸Leu | 1.187 | 0.938 | 1.059 | 0.121 |
| AGU | 丝氨酸Ser | 1.216 | 1.053 | 1.455 | 0.402 |
表3 青钱柳叶绿体基因组最优密码子筛选
Table 3 Optimal codon analysis of the chloroplast genome of Cyclocarya paliurus(Batal.) Iljinsk
| 密码子 Codon | 氨基酸 Amino acid | 相对同义密码子使用度 Relative synonymous codon usage | RSCU低表达 RSCU low expression | RSCU高表达 RSCU high expression | RSCU差值 RSCU difference |
|---|---|---|---|---|---|
| AAU | 天冬酰胺Asn | 1.535 | 1.375 | 1.500 | 0.125 |
| GAA | 谷氨酸Glu | 1.508 | 1.556 | 1.714 | 0.159 |
| CAU | 组氨酸His | 1.574 | 1.000 | 1.200 | 0.200 |
| AAA | 赖氨酸Lys | 1.494 | 1.000 | 1.333 | 0.333 |
| AUU | 异亮氨酸Ile | 1.461 | 1.200 | 1.800 | 0.600 |
| AUG | 蛋氨酸Met | 2.981 | 1.000 | 1.556 | 0.556 |
| GGA | 甘氨酸Gly | 1.634 | 1.647 | 1.846 | 0.199 |
| GGU | 甘氨酸Gly | 1.333 | 0.706 | 0.923 | 0.217 |
| CCU | 脯氨酸Pro | 1.502 | 1.143 | 1.500 | 0.357 |
| ACA | 苏氨酸Thr | 1.213 | 1.091 | 1.500 | 0.409 |
| GUA | 缬氨酸Val | 1.566 | 0.923 | 2.000 | 1.077 |
| GUU | 缬氨酸Val | 1.445 | 1.154 | 1.333 | 0.179 |
| CGU | 精氨酸Arg | 1.304 | 0.278 | 0.545 | 0.268 |
| UUG | 亮氨酸Leu | 1.187 | 0.938 | 1.059 | 0.121 |
| AGU | 丝氨酸Ser | 1.216 | 1.053 | 1.455 | 0.402 |
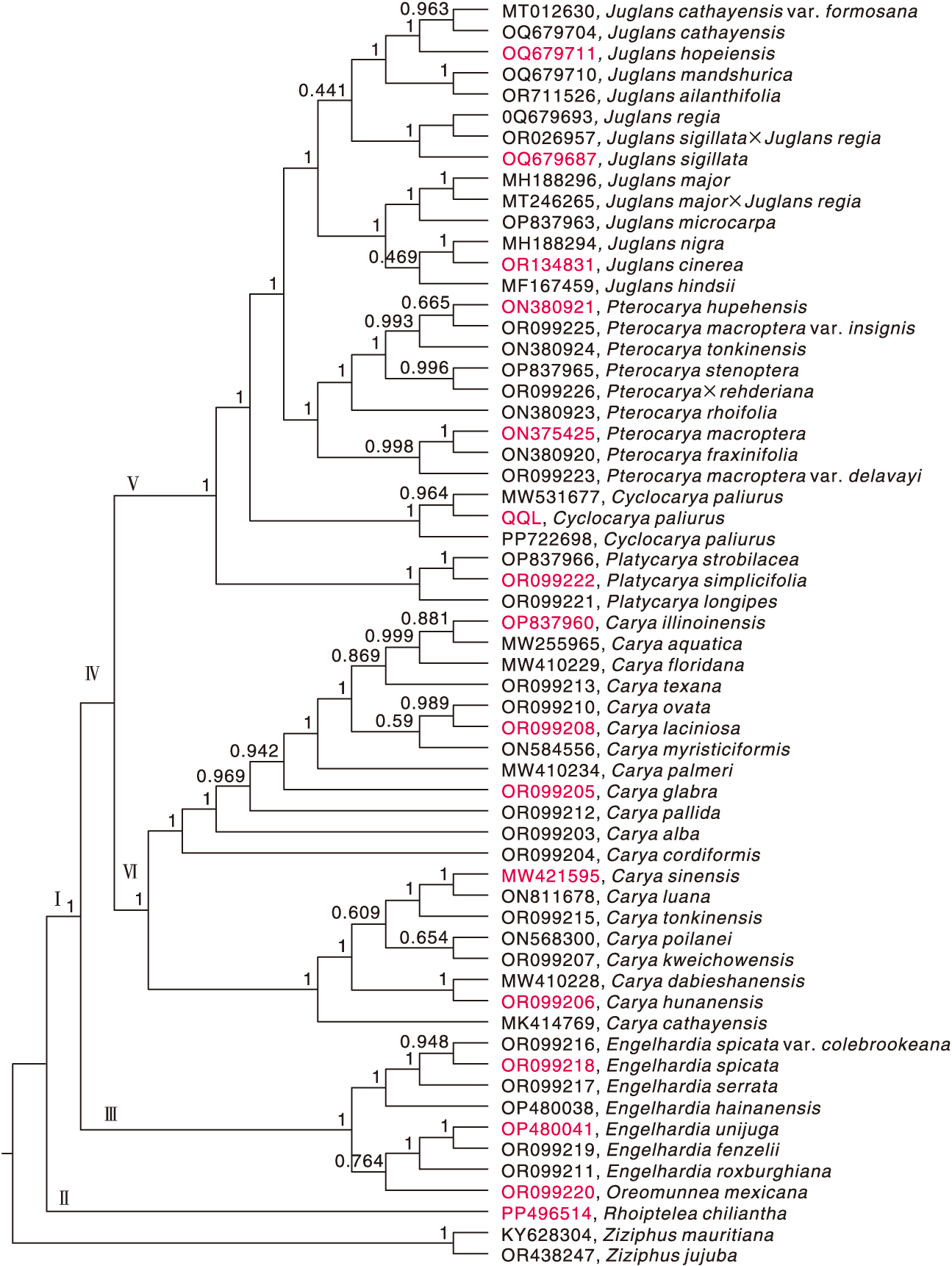
图10 青钱柳及其57个近缘种与2个外群物种叶绿体基因组的系统发育树 红色标注的ID号为用于序列特征、密码子偏好性和系统发育分析的物种,黑色标注的ID号为只用于系统发育分析的物种。
Fig.10 Phylogenetic tree of chloroplast genomes of Cyclocarya paliurus(Batal.) Iljinsk and 57 closely related species, with 2 outgroup species The ID numbers marked in red were species used for sequence features, codon preferences, and phylogenetic analysis, while the ID numbers marked in black were species used only for phylogenetic analysis.
| [1] | 郭春兰, 柳水香, 肖仁才, 等. 我国特有单种属珍稀资源植物青钱柳繁育技术研究进展[J]. 中国野生植物资源, 2024, 43(8): 64-69. |
| GUO C L, LIU S X, XIAO R C, et al. Research progress on breeding technology of Cyclocarya paliurus, a rare plant of single species in China[J]. Chinese Wild Plant Resources, 2024, 43(8): 64-69. | |
| [2] | 刘夏岚, 宋子琪, 胡凤荣, 等. 青钱柳二倍体和四倍体叶特征比较研究[J]. 南京林业大学学报(自然科学版), 2024, 48(4): 76-84. |
| LIU X L, SONG Z Q, HU F R, et al. A comparative study on leaf characters between diploid and tetraploid of Cyclocarya paliurus[J]. Journal of Nanjing Forestry University(Natural Sciences Edition), 2024, 48(4): 76-84. | |
| [3] | 杨杰, 董佳萍, 张露, 等. 青钱柳降糖降尿酸活性部位色谱指纹图谱研究及主成分含量测定[J]. 分析测试学报, 2024, 43(7): 955-962. |
| YANG J, DONG J P, ZHANG L, et al. Chromatographic fingerprinting studies and the main ingredient content determination of hypoglycemic and uric acid-lowering active part of Cyclocarya paliurus[J]. Journal of Instrumental Analysis, 2024, 43(7): 955-962. | |
| [4] | 谷月营, 张赞培, 毛乾兴, 等. 青钱柳无性系对寒驯化的生理响应及其抗寒性初步评价[J]. 生态环境学报, 2024, 33(4): 531-538. |
| GU Y Y, ZHANG Z P, MAO Q X, et al. Physiological response of Cyclocarya paliurus clones to cold acclimation and preliminary evaluation of their cold resistance[J]. Ecology and Environmental Sciences, 2024, 33(4): 531-538. | |
| [5] | 李进, 幸伟年, 何定军, 等. 青钱柳无性系子代测定林早期生长表现[J]. 南方林业科学, 2024, 52(2): 5-9. |
| LI J, XING W N, HE D J, et al. Early growth performance on the clonal progeny test forest of Cyclocarya paliurus[J]. South China Forestry Science, 2024, 52(2): 5-9. | |
| [6] | LI D M, PAN Y G, WU X Y, et al. Comparative chloroplast genomics, phylogenetic relationships and molecular markers development of Aglaonema commutatum and seven green cultivars of Aglaonema[J]. Scientific Reports, 2024, 14: 11820. |
| [7] | CHEN L L, ZHU W, SONG Y, et al. Complete chloroplast genome sequences of endangered tropical Fosbergia species (Family: Rubiaceae)[J]. Forests, 2024, 15(7): 1150. |
| [8] | YIN D B, MU Y T, LI X, et al. Comparison of the complete chloroplast genomes in the Astragalus[J]. Legume Research-an International Journal, 2024: 420-427. |
| [9] | LI B Y, HUANG K, CHEN X L, et al. Comparative and phylogenetic analysis of chloroplast genomes from four species in Quercus section Cyclobalanopsis[J]. BMC Genomic Data, 2024, 25(1): 57. |
| [10] | GAO Z Y, WU Y Z, LI M Z, et al. The auxin response factor (ARF) gene family in Cyclocarya paliurus: genome-wide identification and their expression profiling under heat and drought stresses[J]. Physiology and Molecular Biology of Plants, 2024, 30(6): 921-944. |
| [11] | 黄岩萌, 易炎, 卞国良, 等. 青钱柳染色体的制片技术及核型分析[J]. 东北林业大学学报, 2024, 52(4): 46-51. |
| HUANG Y M, YI Y, BIAN G L, et al. The production technique and karyotype analysis of Cyclocarya paliurus chromosome[J]. Journal of Northeast Forestry University, 2024, 52(4): 46-51. | |
| [12] | 王纪, 方升佐. 不同抗褐化剂对青钱柳愈伤组织酶活性和生长的影响[J]. 南京林业大学学报(自然科学版), 2023, 47(6): 167-174. |
| WANG J, FANG S Z. Effects of different anti-browning agents on enzyme activity and growth in callus of Cyclocarya paliurus[J]. Journal of Nanjing Forestry University(Natural Sciences Edition), 2023, 47(6): 167-174. | |
| [13] | 刘恭孝, 谢磊, 甄学静, 等. 青钱柳发酵物对2型糖尿病合并肝损伤大鼠糖脂代谢及肠道菌群的调节作用[J]. 食品与发酵工业, 2024, 50(12): 220-226. |
| LIU G X, XIE L, ZHEN X J, et al. Exploration of Cyclocarya paliurus fermentation on glucolipid metabolism and intestinal microflora in type 2 diabetes mellitus rats combined with liver injury[J]. Food and Fermentation Industries, 2024, 50(12): 220-226. | |
| [14] | QU Y Q, WANG Q, YU Y H, et al. Comparative analysis of the complete chloroplast genome between tetraploidy and diploidy of Cyclocarya paliurus(Batal.) Iljinskaja[J]. Mitochondrial DNA Part B, Resources, 2021, 6(9): 2669-2671. |
| [15] | HU Y H, YAN J, FENG X J, et al. Characterization of the complete chloroplast genome of wheel wingnut (Cyclocarya paliurus), an endemic in China[J]. Conservation Genetics Resources, 2017, 9(2): 273-275. |
| [16] | PAHLICH E, GERLITZ C. A rapid DNA isolation procedure for small quantities of fresh leaf tissue[J]. Phytochemistry, 1980, 19: 11-13. |
| [17] | LANGDON W B. Performance of genetic programming optimised Bowtie2 on genome comparison and analytic testing (GCAT) benchmarks[J]. BioData Mining, 2015, 8: 1. |
| [18] | DIERCKXSENS N, MARDULYN P, SMITS G. NOVOPlasty: de novo assembly of organelle genomes from whole genome data[J]. Nucleic Acids Research, 2017, 45(4): e18. |
| [19] | HUANG X, MADAN A. CAP3: a DNA sequence assembly program[J]. Genome Research, 1999, 9(9): 868-877. |
| [20] | TILLICH M, LEHWARK P, PELLIZZER T, et al. GeSeq-versatile and accurate annotation of organelle genomes[J]. Nucleic Acids Research, 2017, 45(W1): W6-W11. |
| [21] | LOWE T M, EDDY S R. tRNAscan-SE a program for improved detection of transfer RNA genes in genomic sequence[J]. Nucleic Acids Research, 1997, 25(5): 955-964. |
| [22] | LIU S Y, NI Y, LI J L, et al. CPGView: a package for visualizing detailed chloroplast genome structures[J]. Molecular Ecology Resources, 2023, 23(3): 694-704. |
| [23] | THIEL T, MICHALEK W, VARSHNEY R, et al. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.)[J]. Theoretical and Applied Genetics, 2003, 106(3): 411-422. |
| [24] | KURTZ S, CHOUDHURI J V, OHLEBUSCH E, et al. REPuter: the manifold applications of repeat analysis on a genomic scale[J]. Nucleic Acids Research, 2001, 29(22): 4633-4642. |
| [25] | SHARP P M, LI W H. The codon adaptation index-a measure of directional synonymous codon usage bias, and its potential applications[J]. Nucleic Acids Research, 1987, 15(3): 1281-1295. |
| [26] | KATOH K, STANDLEY D M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability[J]. Molecular Biology and Evolution, 2013, 30(4): 772-780. |
| [27] | AMIRYOUSEFI A, HYVÖNEN J, POCZAI P. IRscope: an online program to visualize the junction sites of chloroplast genomes[J]. Bioinformatics, 2018, 34(17): 3030-3031. |
| [28] | ROZAS J, FERRER-MATA A, SÁNCHEZ-DELBARRIO J C, et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets[J]. Molecular Biology and Evolution, 2017, 34(12): 3299-3302. |
| [29] | 张晶晶, 袁庆, 魏瑶, 等. 忍冬属囊管组植物叶绿体基因组进化分析[J]. 中草药, 2024, 55(9): 3085-3097. |
| ZHANG J J, YUAN Q, WEI Y, et al. Evolutionary analysis of chloroplast genomes of Sect. Isika(Lonicera) species[J]. Chinese Traditional and Herbal Drugs, 2024, 55(9): 3085-3097. | |
| [30] | SUEOKA N. Directional mutation pressure and neutral molecular evolution[J]. Proceedings of the National Academy of Sciences of the United States of America, 1988, 85(8): 2653-2657. |
| [31] | WRIGHT F. The ‘effective number of codons’ used in a gene[J]. Gene, 1990, 87(1): 23-29. |
| [32] | ROMERO H, ZAVALA A, MUSTO H. Codon usage in Chlamydia trachomatis is the result of strand-specific mutational biases and a complex pattern of selective forces[J]. Nucleic Acids Research, 2000, 28(10): 2084-2090. |
| [33] | SUEOKA N. Translation-coupled violation of Parity Rule 2 in human genes is not the cause of heterogeneity of the DNA G+C content of third codon position[J]. Gene, 1999, 238(1): 53-58. |
| [34] | 郭佳星, 黄祥, 杨梅花, 等. 桦木科叶绿体基因组密码子偏好性及系统发育分析[J]. 中国农业科技导报, 2023, 25(10): 74-83. |
| GUOJIA X, HUANG X, YANG M H, et al. Analysis of codon usage bias and phylogenetic in chloroplast genome of Betulaceae[J]. Journal of Agricultural Science and Technology, 2023, 25(10): 74-83. | |
| [35] | LLOYD A T, SHARP P M. Evolution of codon usage patterns: the extent and nature of divergence between Candida albicans and Saccharomyces cerevisiae[J]. Nucleic Acids Research, 1992, 20(20): 5289-5295. |
| [36] | PRICE M N, DEHAL P S, ARKIN A P. FastTree 2: approximately maximum-likelihood trees for large alignments[J]. PLoS One, 2010, 5(3): e9490. |
| [37] | MOHANTA T K, KHAN A L, HASHEM A, et al. Genomic and evolutionary aspects of chloroplast tRNA in monocot plants[J]. BMC Plant Biology, 2019, 19(1): 39. |
| [38] | YAN L J, WANG H L, HUANG X, et al. Chloroplast genomes of genus Tilia: comparative genomics and molecular evolution[J]. Frontiers in Genetics, 2022, 13: 925726. |
| [39] | HU S Q, SHI W B, HUANG Y H, et al. Comparative analysis of complete chloroplast genomes of five Anemone species and phylogenetic analysis within Tribe Anemoneae(Ranunculaceae)[J]. Journal of Plant Biochemistry and Biotechnology, 2024, 33(3): 271-287. |
| [40] | QU K, CHEN Y, LIU D, et al. Comprehensive analysis of the complete mitochondrial genome of Lilium tsingtauense reveals a novel multichromosome structure[J]. Plant Cell Reports, 2024, 43(6): 150. |
| [41] | PEI D, YU X, FU W H, et al. The evolution and formation of centromeric repeats analysis in Vitis vinifera[J]. Planta, 2024, 259(5): 99. |
| [42] | HAN F C, BI C W, ZHAO Y X, et al. Unraveling the complex evolutionary features of the Cinnamomum camphora mitochondrial genome[J]. Plant Cell Reports, 2024, 43(7): 183. |
| [43] | XU S J, TENG K, ZHANG H, et al. Chloroplast genomes of four Carex species: long repetitive sequences trigger dramatic changes in chloroplast genome structure[J]. Frontiers in Plant Science, 2023, 14: 1100876. |
| [44] | LI L, HU Y F, HE M, et al. Comparative chloroplast genomes: insights into the evolution of the chloroplast genome of Camellia sinensis and the phylogeny of Camellia[J]. BMC Genomics, 2021, 22(1): 138. |
| [45] | CHEN S Y, LI K, CAO W Q, et al. Codon-resolution analysis reveals a direct and context-dependent impact of individual synonymous mutations on mRNA level[J]. Molecular Biology and Evolution, 2017, 34(11): 2944-2958. |
| [46] | 罗秦欢, 段炼, 赵月梅. “红阳”猕猴桃叶绿体基因组密码子使用偏性分析[J]. 西南林业大学学报(自然科学), 2024, 44(6): 81-88. |
| LUO Q H, DUAN L, ZHAO Y M. Codon usage bias of chloroplast genome in Actinidia chinensis ‘Hongyang’[J]. Journal of Southwest Forestry University(Natural Sciences), 2024, 44(6): 81-88. | |
| [47] | WALIA G K, DHILLON G K. DNA barcoding and molecular phylogeny of genus Pseudagrion(Odonata: Coenagrionidae) based on mitochondrial COI ND1 and 16S rRNA genes[J]. International Journal of Tropical Insect Science, 2024, 44(4): 1567-1590. |
| [48] | WANG H J, WU Z G, LI T, et al. Highly active repeat-mediated recombination in the mitogenome of the aquatic grass Hygroryza aristata[J]. BMC Plant Biology, 2024, 24(1): 644. |
| [49] | NOROOZI M, GHAHREMANINEJAD F, RIAHI M, et al. Phylogenomics and plastome evolution of lithospermeae (Boraginaceae)[J]. BMC Plant Biology, 2024, 24(1): 957. |
| [50] | DELSUC F, BRINKMANN H, PHILIPPE H. Phylogenomics and the reconstruction of the tree of life[J]. Nature Reviews Genetics, 2005, 6(5): 361-375. |
| [51] | QU Y Q, SHANG X L, ZENG Z Y, et al. Whole-genome duplication reshaped adaptive evolution in a relict plant species, Cyclocarya paliurus[J]. Genomics, Proteomics & Bioinformatics, 2023, 21(3): 455-469. |
| [52] | 田慧, 陈宏夏, 黄勇, 等. 青钱柳种质资源SCoT分子标记遗传多样性分析[J]. 世界科学技术-中医药现代化, 2022, 24(7): 2732-2739. |
| TIAN H, CHEN H X, HUANG Y, et al. Genetic diversity of germplasm resource SCoT of Cyclocarya paliurus(Batal.) Iljinsk by molecular markers[J]. World Science and Technology-Modernization of Traditional Chinese Medicine, 2022, 24(7): 2732-2739. | |
| [53] | 陈秀娟, 柏明娥, 王丽玲, 等. 青钱柳种质资源亲缘关系的ISSR分析评价[J]. 中国林副特产, 2016(4): 6-10. |
| CHEN X J, BAI M E, WANG L L, et al. ISSR analysis and evaluation of genetic relationship of Cyclocarya paliurus germplasm resources[J]. Forest By-Product and Speciality in China, 2016(4): 6-10. |
| [1] | 张加强, 刘慧春, 王杰, 许雯婷, 周江华, 朱开元. 金银花大毛花叶绿体基因组密码子的偏好性分析[J]. 浙江农业学报, 2023, 35(4): 821-830. |
| [2] | 楚志刚, 田云芳. 蕙兰一个PEBP家族基因的克隆及生物信息学分析[J]. 浙江农业学报, 2022, 34(8): 1679-1691. |
| [3] | 蒋瑞平, 赵辰晖, 李文杰, 安秋菊, 李佳伦, 周嘉裕, 李遂焰, 廖海. 豆科植物IPI基因密码子偏好性[J]. 浙江农业学报, 2022, 34(6): 1114-1123. |
| [4] | 韦海忠, 潘丽芹, 汤紫依, 田盛野, 何海叶, 尹龙飞, 郑德伟, 张慧娟, 蒋明. 青花菜SKIP基因的克隆与表达分析[J]. 浙江农业学报, 2022, 34(5): 966-973. |
| [5] | 梁丽琴, 杨瑞, 郜刚. 马铃薯StUOXs基因家族的生物信息学分析[J]. 浙江农业学报, 2020, 32(9): 1523-1532. |
| [6] | 宗秋芳, 黄焱杰, 吴丽思, 吴圣龙, 包文斌. 猪Claudin家族基因密码子使用偏好性分析[J]. 浙江农业学报, 2018, 30(12): 2007-2017. |
| [7] | 冯琛, 汤浩茹, 江雷雨, 宋霞, 张云婷, 叶云天, 陈清, 孙勃. 红蓝光对草莓转录组特异表达基因密码子使用偏好性的影响[J]. 浙江农业学报, 2017, 29(4): 566-574. |
| [8] | 邹莉,杨苑艺,孙婷婷,王世新,崔嵘,李晶莹. 野生花脸香蘑的分离纯化及ITS序列鉴定[J]. 浙江农业学报, 2016, 28(2): 264-. |
| [9] | 李春,褚威威,刘欢,费文超,韩苗苗. 基于拟氨基酸多重集的DNA序列的数值刻画及其应用[J]. 浙江农业学报, 2015, 27(7): 1244-. |
| [10] | 李喜莲,杨元杰,李倩,刘金殿,顾志敏*. 螯虾次目功能基因密码子偏好性研究[J]. 浙江农业学报, 2014, 26(4): 862-. |
| [11] | 肖竞;孙建义;周德平;陈艳;付亮剑. 一株产生内切β-1,4-木聚糖酶的菌株的分离、鉴定及其酶学特性研究[J]. , 2003, 15(2): 0-61. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||