浙江农业学报 ›› 2026, Vol. 38 ›› Issue (2): 269-283.DOI: 10.3969/j.issn.1004-1524.20241050
膜荚黄芪TCP基因家族全基因组分析及其对非生物胁迫的响应
a.浙江中医药大学 第一临床医学院 浙江 杭州 310053 b.浙江中医药大学 基础医学院 浙江 杭州 310053
-
收稿日期:2024-12-02出版日期:2026-02-25发布日期:2026-03-24 -
作者简介:马琦洋,主要从事中药资源鉴定与草药分子生理研究。E-mail: mqy20242025@163.com -
通讯作者:*黄雪莲,E-mail: h25872968110xl@126.com -
基金资助:省级创业训练计划(S202310344003X);2023浙江省新苗人才计划(2023R410004)
Genome-wide analysis of the TCP gene family in Astragalus membranaceus (Fisch.) Bunge and its response to abiotic stress
MA Qiyanga( ), HUANG Xuelianb,*(
), HUANG Xuelianb,*( )
)
a. The First Clinical Medical College, Zhejiang Chinese Medical University Hangzhou 310053, China b. School of Basic Medical Sciences ,Zhejiang Chinese Medical University Hangzhou 310053, China
-
Received:2024-12-02Published:2026-02-25Online:2026-03-24
摘要:
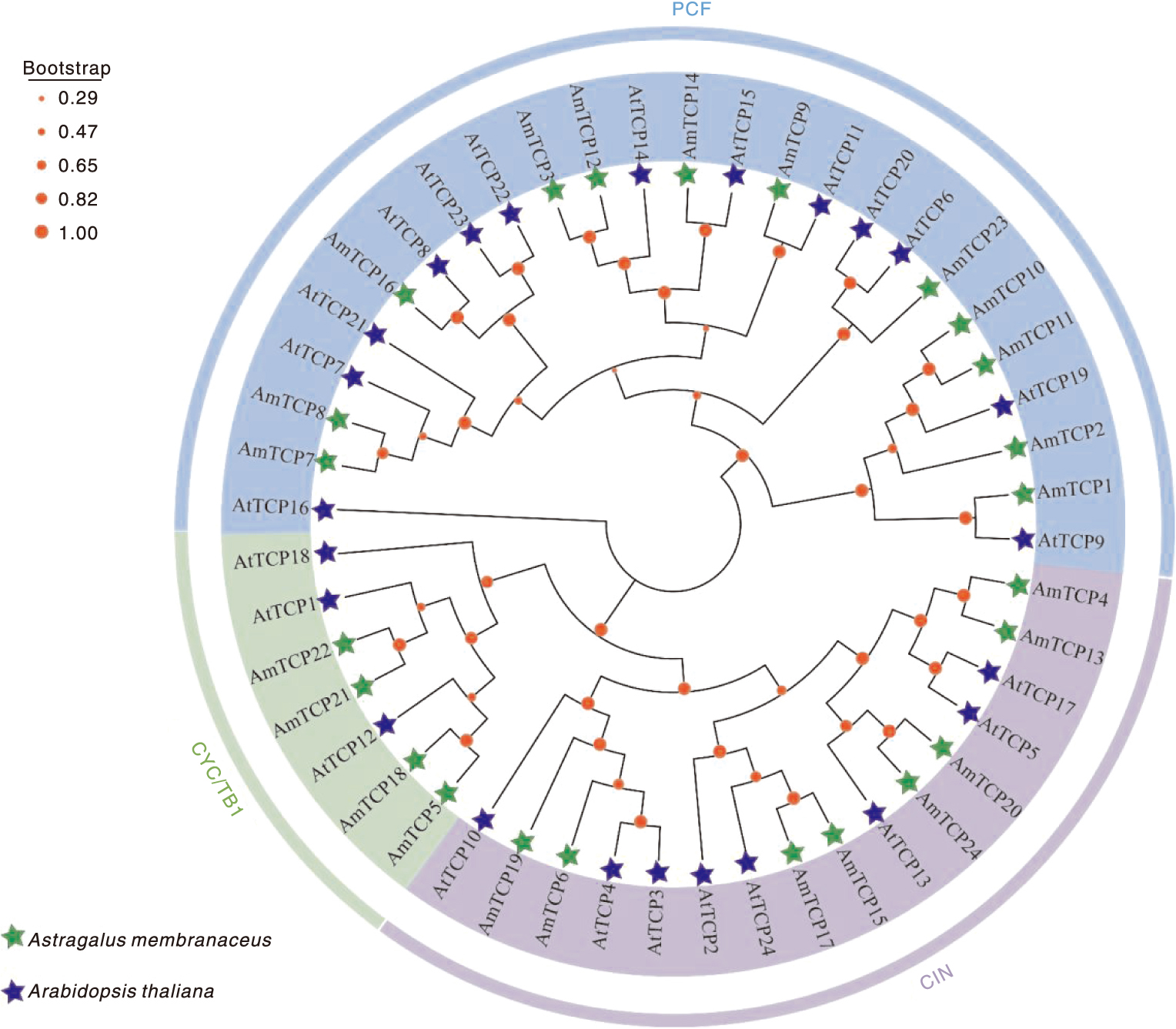
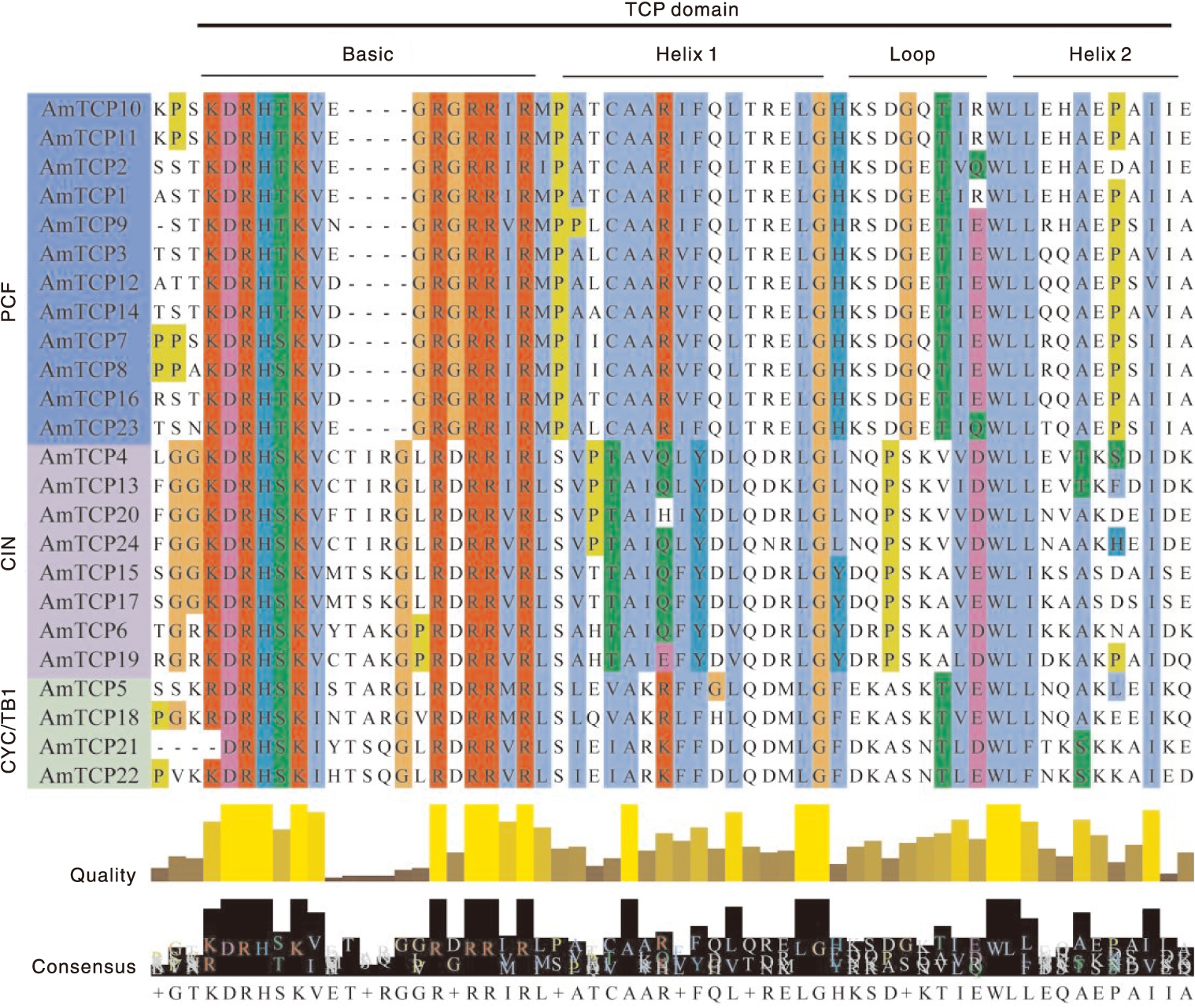
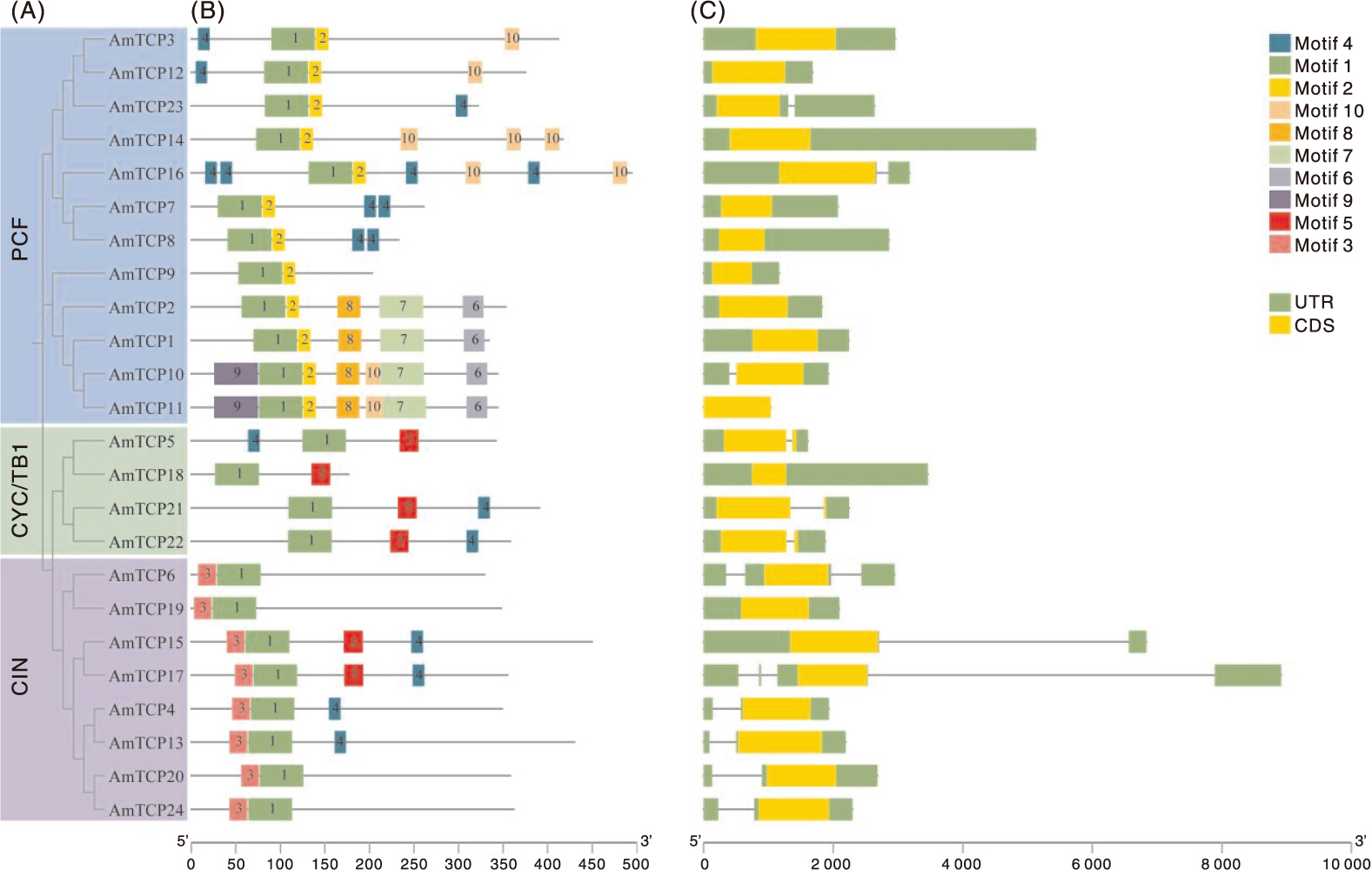
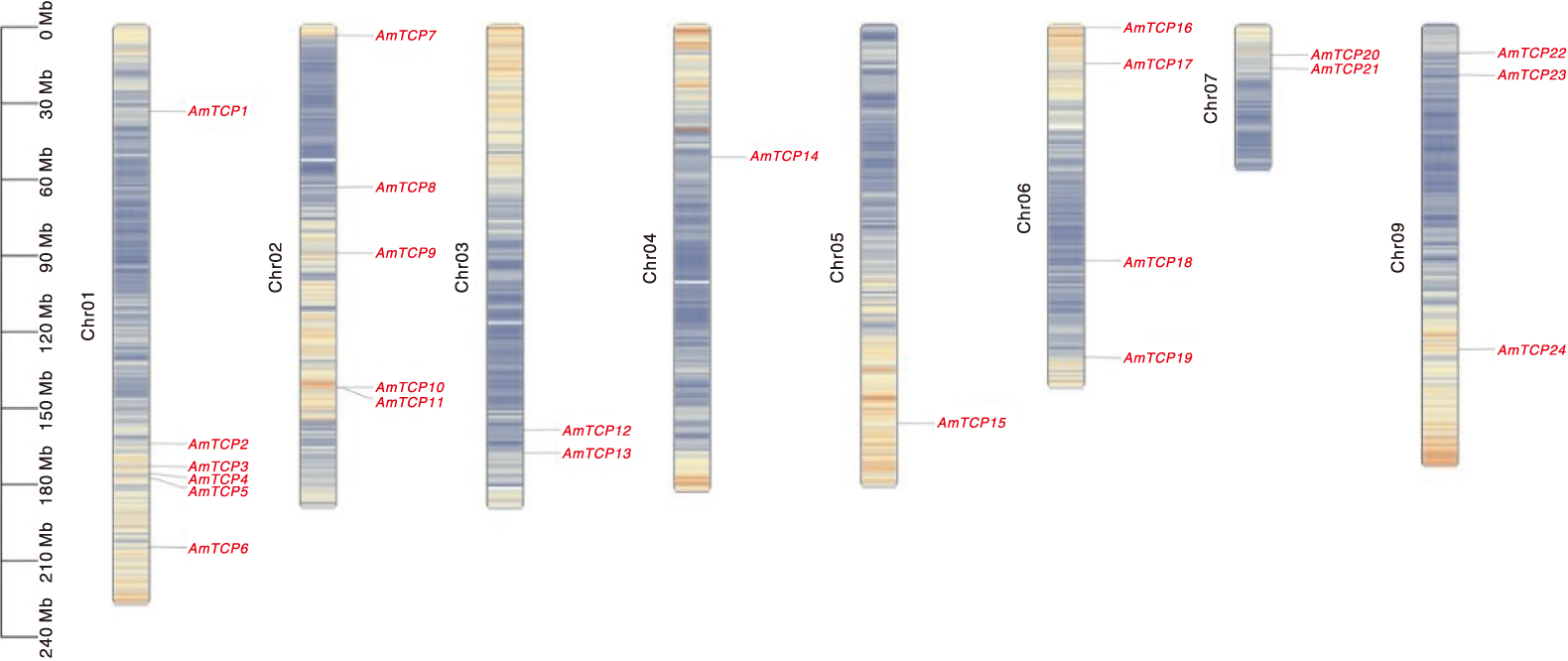
本研究旨在鉴定和分析膜荚黄芪TCP基因家族,并揭示其对非生物胁迫的响应,为研究膜荚黄芪TCP基因在非生物胁迫中的功能提供理论依据。通过生物信息学方法对膜荚黄芪TCP基因及其编码的氨基酸序列进行系统分析,利用RNA-seq技术研究膜荚黄芪TCP基因在不同组织(根、茎和叶)中的表达模式,并通过实时荧光定量PCR(RT-qPCR)验证了5个代表性AmTCP基因在非生物胁迫(高温、干旱、盐胁迫)条件下的表达谱。结果表明,共鉴定出24个TCP基因且不均匀分布于膜荚黄芪的8条染色体上。24个AmTCP基因可分为Ⅰ类(PCF)和Ⅱ类(CYC/TB1和CIN)。同一亚家族蛋白具有高度相似的基序组成,大多数AmTCP基因缺乏内含子。片段重复在膜荚黄芪TCP基因家族进化中发挥了重要作用。此外,启动子区域分析发现多个光响应元件和激素响应元件。组织表达分析显示,AmTCP基因具有显著的组织表达特异性。在非生物胁迫下,5个代表性AmTCP基因的表达水平存在显著差异,其中AmTCP18和AmTCP19基因在多种胁迫条件下均表现出较高的表达水平。综上所述,AmTCP18和AmTCP19可能在膜荚黄芪对非生物胁迫的响应中发挥关键作用,本研究为膜荚黄芪TCP基因家族的进一步功能和进化研究奠定了基础。
中图分类号:
引用本文
马琦洋, 黄雪莲. 膜荚黄芪TCP基因家族全基因组分析及其对非生物胁迫的响应[J]. 浙江农业学报, 2026, 38(2): 269-283.
MA Qiyang, HUANG Xuelian. Genome-wide analysis of the TCP gene family in Astragalus membranaceus (Fisch.) Bunge and its response to abiotic stress[J]. Acta Agriculturae Zhejiangensis, 2026, 38(2): 269-283.
| 基因 Gene | 正向引物(5'→3') Forward primer (5'→3') | 反向引物(5'→3') Reverse primer (5'→3') |
|---|---|---|
| Actin | CTCTCTCAGCACCTTCGAGCAG | TCCACATACAACCGCTCCACTG |
| AmTCP8 | CCAGGCGGAACCGAGCATTATTG | TCGCTTCCTGCTGCTGATGATGA |
| AmTCP16 | AACGGCGATGATGGCGATGATG | TGCTGCTGCTGCTGCTGATG |
| AmTCP18 | GCGGCAGCGGCAACTATAACAA | GCTCGGAATATGATGATGATGCGGAAT |
| AmTCP19 | AGCAGCAGCAGCAGCAGAAC | CGTTGTTGCCGGTATCCAGGTT |
| AmTCP23 | AGCCGCACCAACTGGAACATTAC | TCTGATGCTGATGCTGGTTCTGATG |
表1 RT-qPCR引物
Table 1 Primers for RT-qPCR
| 基因 Gene | 正向引物(5'→3') Forward primer (5'→3') | 反向引物(5'→3') Reverse primer (5'→3') |
|---|---|---|
| Actin | CTCTCTCAGCACCTTCGAGCAG | TCCACATACAACCGCTCCACTG |
| AmTCP8 | CCAGGCGGAACCGAGCATTATTG | TCGCTTCCTGCTGCTGATGATGA |
| AmTCP16 | AACGGCGATGATGGCGATGATG | TGCTGCTGCTGCTGCTGATG |
| AmTCP18 | GCGGCAGCGGCAACTATAACAA | GCTCGGAATATGATGATGATGCGGAAT |
| AmTCP19 | AGCAGCAGCAGCAGCAGAAC | CGTTGTTGCCGGTATCCAGGTT |
| AmTCP23 | AGCCGCACCAACTGGAACATTAC | TCTGATGCTGATGCTGGTTCTGATG |
| 蛋白名 Protein name | 基因名 Gene name | 氨基酸数量 Amino acid quantity | 相对分子质量/ku Relative molecular weight/ku | 等电点 Isoelectric point | 不稳定系数 Instability index | 脂肪指数 Aliphatic index | 亲水性 Hydrophilicity |
|---|---|---|---|---|---|---|---|
| AmTCP1 | Amem01G08650.1 | 334 | 36.25 | 9.62 | 61.72 | 72.78 | -0.355 |
| AmTCP2 | Amem01G26990.1 | 353 | 37.44 | 5.99 | 52.15 | 67.73 | -0.460 |
| AmTCP3 | Amem01G29700.1 | 412 | 44.15 | 7.97 | 67.32 | 58.33 | -0.720 |
| AmTCP4 | Amem01G30870.1 | 349 | 39.08 | 8.44 | 53.65 | 55.62 | -0.756 |
| AmTCP5 | Amem01G31480.1 | 342 | 39.30 | 9.16 | 57.79 | 55.85 | -1.099 |
| AmTCP6 | Amem01G39660.1 | 329 | 35.68 | 6.21 | 43.87 | 57.63 | -0.681 |
| AmTCP7 | Amem02G01750.1 | 261 | 27.81 | 9.56 | 48.82 | 70.73 | -0.475 |
| AmTCP8 | Amem02G07350.1 | 233 | 25.13 | 9.10 | 52.77 | 70.52 | -0.566 |
| AmTCP9 | Amem02G13390.1 | 203 | 21.44 | 8.44 | 61.98 | 68.82 | -0.265 |
| AmTCP10 | Amem02G30910.1 | 344 | 36.12 | 5.46 | 54.31 | 65.00 | -0.358 |
| AmTCP11 | Amem02G31020.1 | 344 | 36.13 | 5.46 | 53.03 | 65.00 | -0.358 |
| AmTCP12 | Amem03G33990.1 | 375 | 41.03 | 6.90 | 66.54 | 52.88 | -0.784 |
| AmTCP13 | Amem03G35070.1 | 430 | 47.97 | 7.42 | 47.37 | 59.35 | -0.770 |
| AmTCP14 | Amem04G15690.1 | 417 | 44.13 | 6.90 | 56.30 | 57.15 | -0.661 |
| AmTCP15 | Amem05G34130.3 | 450 | 49.85 | 7.21 | 52.74 | 52.27 | -0.986 |
| AmTCP16 | Amem06G00490.1 | 494 | 52.19 | 6.15 | 62.09 | 50.63 | -0.856 |
| AmTCP17 | Amem06G06270.2 | 355 | 39.83 | 8.56 | 58.10 | 55.55 | -1.028 |
| AmTCP18 | Amem06G19960.1 | 177 | 20.64 | 10.50 | 49.17 | 69.44 | -0.921 |
| AmTCP19 | Amem06G25360.1 | 348 | 37.92 | 5.67 | 57.14 | 59.17 | -0.719 |
| AmTCP20 | Amem07G03630.1 | 358 | 40.03 | 8.28 | 49.00 | 71.84 | -0.665 |
| AmTCP21 | Amem07G04880.1 | 391 | 44.30 | 9.12 | 51.58 | 66.14 | -0.874 |
| AmTCP22 | Amem09G02360.1 | 358 | 40.20 | 8.26 | 54.21 | 55.89 | -0.885 |
| AmTCP23 | Amem09G03560.1 | 322 | 35.47 | 9.62 | 52.23 | 58.79 | -0.730 |
| AmTCP24 | Amem09G20040.1 | 362 | 40.59 | 8.87 | 44.80 | 69.53 | -0.735 |
表2 24个AmTCP基因及其编码蛋白的理化性质
Table 2 Physicochemical characteristics of 24 AmTCP genes and their encoded proteins
| 蛋白名 Protein name | 基因名 Gene name | 氨基酸数量 Amino acid quantity | 相对分子质量/ku Relative molecular weight/ku | 等电点 Isoelectric point | 不稳定系数 Instability index | 脂肪指数 Aliphatic index | 亲水性 Hydrophilicity |
|---|---|---|---|---|---|---|---|
| AmTCP1 | Amem01G08650.1 | 334 | 36.25 | 9.62 | 61.72 | 72.78 | -0.355 |
| AmTCP2 | Amem01G26990.1 | 353 | 37.44 | 5.99 | 52.15 | 67.73 | -0.460 |
| AmTCP3 | Amem01G29700.1 | 412 | 44.15 | 7.97 | 67.32 | 58.33 | -0.720 |
| AmTCP4 | Amem01G30870.1 | 349 | 39.08 | 8.44 | 53.65 | 55.62 | -0.756 |
| AmTCP5 | Amem01G31480.1 | 342 | 39.30 | 9.16 | 57.79 | 55.85 | -1.099 |
| AmTCP6 | Amem01G39660.1 | 329 | 35.68 | 6.21 | 43.87 | 57.63 | -0.681 |
| AmTCP7 | Amem02G01750.1 | 261 | 27.81 | 9.56 | 48.82 | 70.73 | -0.475 |
| AmTCP8 | Amem02G07350.1 | 233 | 25.13 | 9.10 | 52.77 | 70.52 | -0.566 |
| AmTCP9 | Amem02G13390.1 | 203 | 21.44 | 8.44 | 61.98 | 68.82 | -0.265 |
| AmTCP10 | Amem02G30910.1 | 344 | 36.12 | 5.46 | 54.31 | 65.00 | -0.358 |
| AmTCP11 | Amem02G31020.1 | 344 | 36.13 | 5.46 | 53.03 | 65.00 | -0.358 |
| AmTCP12 | Amem03G33990.1 | 375 | 41.03 | 6.90 | 66.54 | 52.88 | -0.784 |
| AmTCP13 | Amem03G35070.1 | 430 | 47.97 | 7.42 | 47.37 | 59.35 | -0.770 |
| AmTCP14 | Amem04G15690.1 | 417 | 44.13 | 6.90 | 56.30 | 57.15 | -0.661 |
| AmTCP15 | Amem05G34130.3 | 450 | 49.85 | 7.21 | 52.74 | 52.27 | -0.986 |
| AmTCP16 | Amem06G00490.1 | 494 | 52.19 | 6.15 | 62.09 | 50.63 | -0.856 |
| AmTCP17 | Amem06G06270.2 | 355 | 39.83 | 8.56 | 58.10 | 55.55 | -1.028 |
| AmTCP18 | Amem06G19960.1 | 177 | 20.64 | 10.50 | 49.17 | 69.44 | -0.921 |
| AmTCP19 | Amem06G25360.1 | 348 | 37.92 | 5.67 | 57.14 | 59.17 | -0.719 |
| AmTCP20 | Amem07G03630.1 | 358 | 40.03 | 8.28 | 49.00 | 71.84 | -0.665 |
| AmTCP21 | Amem07G04880.1 | 391 | 44.30 | 9.12 | 51.58 | 66.14 | -0.874 |
| AmTCP22 | Amem09G02360.1 | 358 | 40.20 | 8.26 | 54.21 | 55.89 | -0.885 |
| AmTCP23 | Amem09G03560.1 | 322 | 35.47 | 9.62 | 52.23 | 58.79 | -0.730 |
| AmTCP24 | Amem09G20040.1 | 362 | 40.59 | 8.87 | 44.80 | 69.53 | -0.735 |
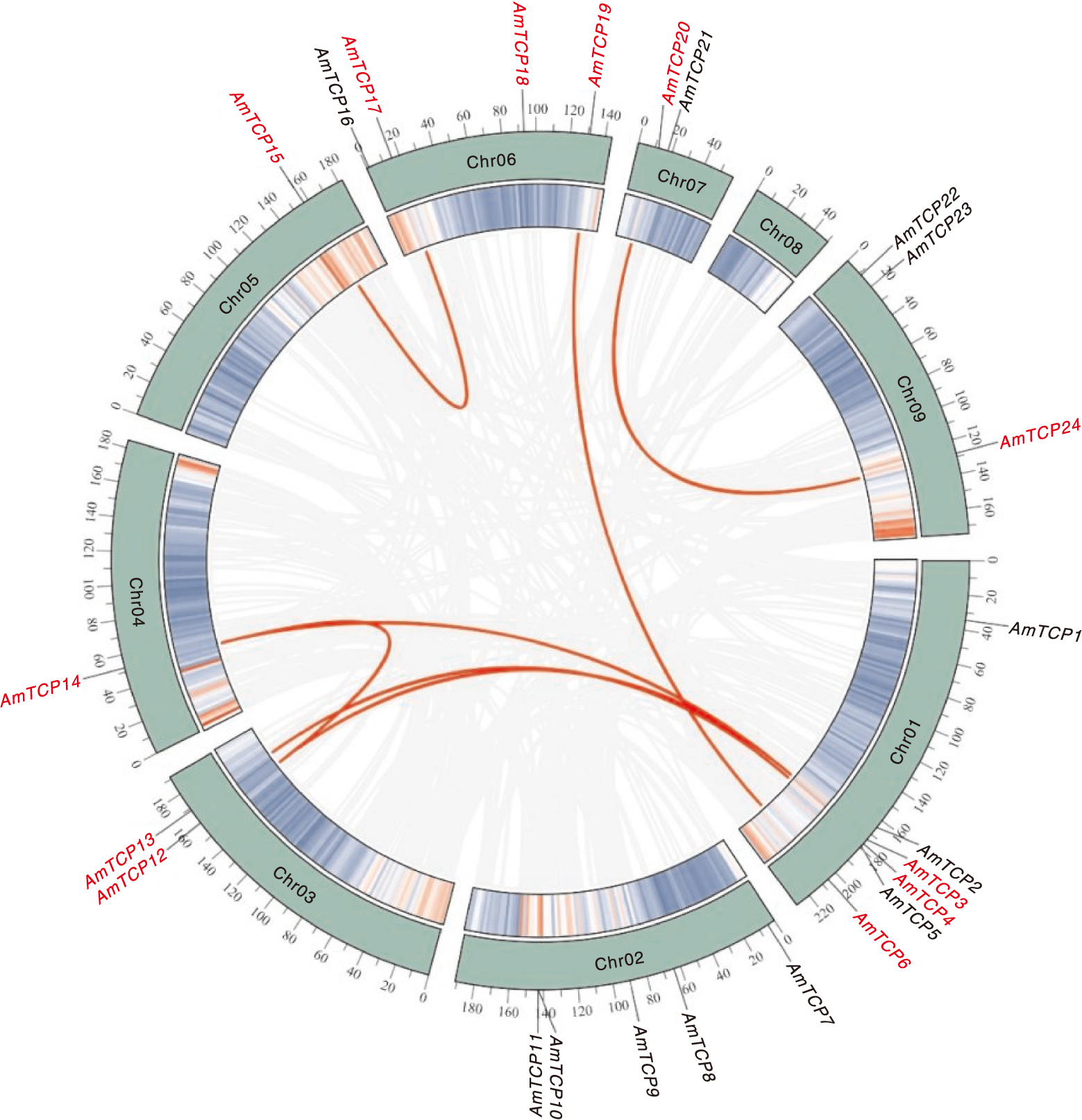
图5 AmTCP基因的共线性 内环的绿线代表每条染色体的基因密度,共线性基因用浅灰色线表示,AmTCP的种内共线性基因对用红色曲线表示,字体也标为红色。
Fig.5 Colinearity of the AmTCP gene The green line in the inner ring represents the gene density per chromosome, co-lined genes are indicated by light gray lines, and intraspecific co-lined gene pairs of AmTCP are labeled with red curves and the font is also labeled in red.
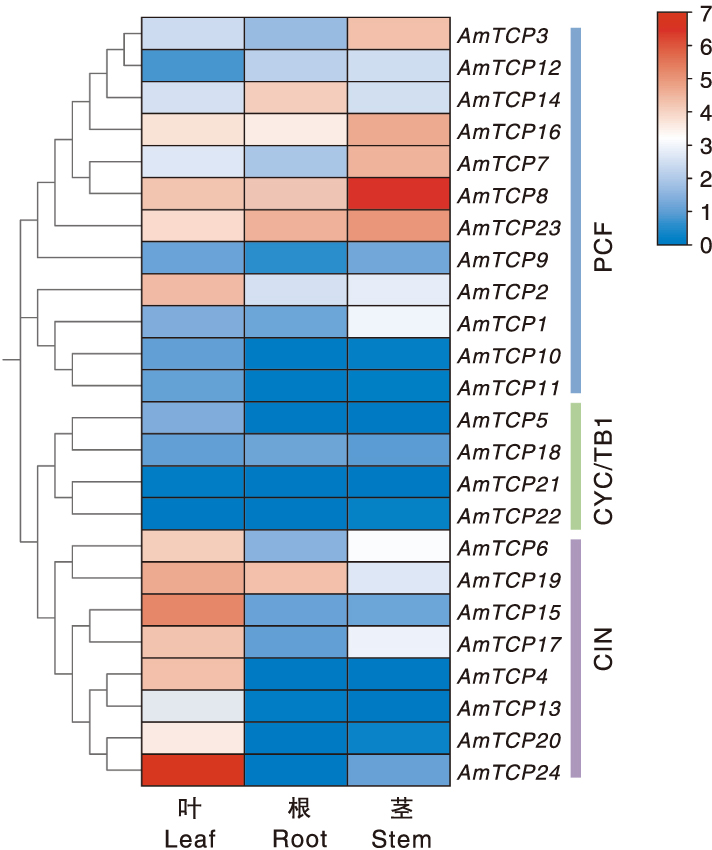
图8 AmTCP基因在不同组织的表达 颜色条代表标准化表达水平(log2转换的倍数变化)。红色和蓝色表示上调和下调的基因,白色表示无表达。右边蓝色、绿色和紫色条带分别表示PCF、CYC/TB1和CIN亚家族。
Fig.8 AmTCP gene expression in different tissues Color bars represent normalised expression levels (log2 converted fold change). Red and blue indicate up- and down-regulated genes, and white indicates no expression. The blue, green and purple bands on the right represent the PCF, CYC/TB1 and CIN subfamilies.
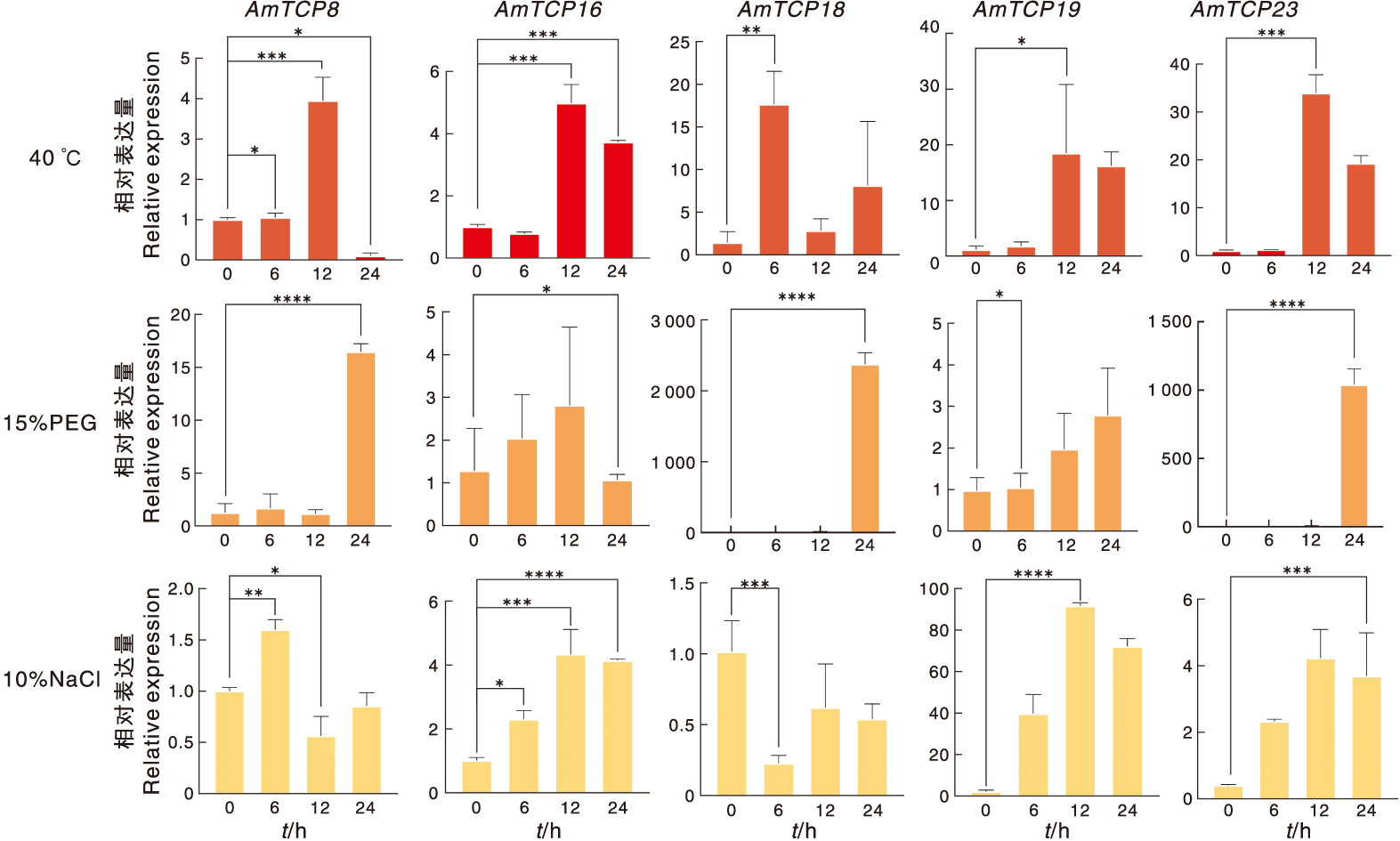
图9 AmTCP基因在非生物胁迫下的表达 高温(40 ℃),干旱(15% PEG),盐(10% NaCl)。“*”表示不同处理间在0.05水平差异显著,“**”表示不同处理间在0.01水平差异显著,“***”表示不同处理间在0.000 5水平差异显著,“****”表示不同处理间在0.000 1水平差异显著。
Fig.9 Expression of AmTCP genes under abiotic stress High temperature (40 ℃), drought (15% PEG), salt (10% NaCl). “*” indicates a significant difference at the 0.05 level between different treatments, “**” indicates a significant difference at the 0.01 level between different treatments, “***” indicates a significant difference at the 0.000 5 level between different treatments, “****” indicates a significant difference at the 0.000 1 level between different treatments.
| [1] | CUBAS P, LAUTER N, DOEBLEY J, et al. The TCP domain: a motif found in proteins regulating plant growth and development[J]. The Plant Journal, 1999, 18(2): 215-222. |
| [2] | DOEBLEY J, STEC A, HUBBARD L. The evolution of apical dominance in maize[J]. Nature, 1997, 386(6624): 485-488. |
| [3] | LUO D, CARPENTER R, VINCENT C, et al. Origin of floral asymmetry in Antirrhinum[J]. Nature, 1996, 383(6603): 794-799. |
| [4] | KOSUGI S, OHASHI Y. PCF1 and PCF2 specifically bind to cis elements in the rice proliferating cell nuclear antigen gene[J]. The Plant Cell, 1997, 9(9): 1607-1619. |
| [5] | AGGARWAL P, DAS GUPTA M, JOSEPH A P, et al. Identification of specific DNA binding residues in the TCP family of transcription factors in Arabidopsis[J]. The Plant Cell, 2010, 22(4): 1174-1189. |
| [6] | MARTÍN-TRILLO M, CUBAS P. TCP genes: a family snapshot ten years later[J]. Trends in Plant Science, 2010, 15(1): 31-39. |
| [7] | PARAPUNOVA V, BUSSCHER M, BUSSCHER-LANGE J, et al. Identification, cloning and characterization of the tomato TCP transcription factor family[J]. BMC Plant Biology, 2014, 14(1): 157. |
| [8] | LI Z, LI B, SHEN W, et al. Comprehensive genomic identification and expression analysis of TCP transcription factors in kiwifruit (Actinidia chinensis)[J]. BMC Genomics, 2019, 19(1): 271. |
| [9] | QI X H, SUN S L, XU P, et al. Genome-wide identification and expression analysis of the TCP transcription factor genes in Capsicum annuum L.[J]. International Journal of Molecular Sciences, 2016, 17(6): 702. |
| [10] | WANG Y, CHEN D, WANG D, et al. Genome-wide analysis of TCP transcription factor gene family in Medicago sativa reveals significant roles in shoot lateral organ development[J]. BMC Plant Biology, 2015, 15: 29. |
| [11] | LIU C, LYU T T, SHEN Y H, et al. Genome-wide identification and integrated analysis of TCP genes controlling ginsenoside biosynthesis in Panax ginseng[J]. BMC Plant Biology, 2024, 24(1): 47. |
| [12] | KOYAMA T, SATO F, OHME-TAKAGI M. Roles of miR319 and TCP transcription factors in leaf development[J]. Plant Physiology, 2017, 175(2): 874-885. |
| [13] | GUAN P, RIPOLL J J, WANG R, et al. Interacting TCP and NLP transcription factors control plant responses to nitrate availability[J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(9): 2419-2424. |
| [14] | MUKHOPADHYAY P, TYAGI A K. OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways[J]. Scientific Reports, 2015, 5: 9998. |
| [15] | SHARMA R, SINGH G, BHATTACHARYA S, et al. A genome-wide survey of TCP transcription factors provides insights into their functional roles in chickpea (Cicer arietinum L.)[J]. Scientific Reports, 2018, 8(1): 8455. |
| [16] | WU X Y, LI J Z, WEN X H, et al. Genome-wide identification of the TCP gene family in Chrysanthemum lavandulifolium and its homologs expression patterns during flower development in different Chrysanthemum species[J]. Frontiers in Plant Science, 2023, 14(1): 1276123. |
| [17] | CHAI W, JIANG P, HUANG G, et al. ZmTCP42, a TCP transcription factor from maize, confers drought tolerance in transgenic Arabidopsis[J]. International Journal of Molecular Sciences, 2021, 22(4): 2024. |
| [18] | MA J, WANG Q L, SUN R R, et al. Genome-wide identification and expression analysis of TCP transcription factors in Gossypium raimondii[J]. Scientific Reports, 2014, 4: 6645. |
| [19] | 后叶虎, 邱璐, 靳晓杰, 等. 基于化学生物信息学方法探讨黄芪通过调节能量代谢发挥“补气”功效的物质基础和分子机制[J]. 中国现代应用药学, 2023, 40(14): 1906. |
| HOU Y H, QIU L, JIN X J, et al. Exploration on material basis and molecular mechanism of astragali radix exerting the effect of “Qi-invigorating” through regulating energy metabolism based on chemo-bio informatics methods[J]. Chinese Journal of Modern Applied Pharmacy, 2023, 40(14): 1906. | |
| [20] | 余意, 胡明华, 张丹丹, 等. 黄芪多糖对气虚大鼠的补气作用及其机制探讨[J]. 中药新药与临床药理, 2021, 32(4): 505. |
| YU Y, HU M H, ZHANG D D, et al. The “Qi-invigorating” effect and possible underlying mechanism of astragalus polysaccharides on rats with Qi-deficiency[J]. Chinese Journal of New Drugs and Clinical Remedies, 2021, 32(4): 505. | |
| [21] | 周文静, 柴智, 刘佳欣, 等. 黄芪补气功用探微[J]. 中国民间疗法, 2021, 29(20): 1. |
| ZHOU W J, CHAI Z, LIU J X, et al. Study on the function of “Qi-invigorating” with Astragalus membranaceus[J]. Chinese Folk Therapy, 2021, 29(20): 1. | |
| [22] | 张倩倩, 李光跃, 苏优拉, 等. 干旱胁迫对蒙古黄芪和膜荚黄芪不同器官黄酮类成分积累的影响[J]. 西北植物学报, 2020, 40(7): 1201. |
| ZHANG Q Q, LI G Y, SU Y L, et al. Effects of drought stress on flavonoid accumulation in different organs of Astragalus mongholicus and Astragalus membranaceus[J]. Acta Botanica Boreali-Occidentalia Sinica, 2020, 40(7): 1201. | |
| [23] | 王楠, 高静, 岳正刚, 等. 黄芪种子萌发对光温条件和旱盐胁迫的响应特征[J]. 草业学报, 2018, 27(1): 106. |
| WANG N, GAO J, YUE Z G, et al. Response characteristics of Astragalus seed germination to light-temperature conditions and drought-salt stress[J]. Acta Prataculturae Sinica, 2018, 27(1): 106. | |
| [24] | 刘洋. 应用代谢组学对比研究两种黄芪响应UV-B和干旱胁迫的代谢基础[D]. 哈尔滨: 东北林业大学, 2016. |
| LIU Y. Comparative study on the metabolic basis of two Astragalus membranaceus species in response to UV-B and drought stress based on metabolomics[D]. Harbin: Northeast Forestry University, 2016. | |
| [25] | WANG J L, WANG H W, CAO Y N, et al. Comprehensive evolutionary analysis of the TCP gene family: Further insights for its origin, expansion, and diversification[J]. Frontiers in Plant Science, 2022, 13: 994567. |
| [26] | ZHOU H Y, HWARARI D, MA H Y, et al. Genomic survey of TCP transcription factors in plants: phylogenomics, evolution and their biology[J]. Frontiers in Genetics, 2022, 13: 1060546. |
| [27] | FAN H J, CHAI Z, YANG X K, et al. Chromosome-scale genome assembly of Astragalus membranaceus using PacBio and Hi-C technologies[J]. Scientific Data, 2024, 11(1):1071. |
| [28] | CHEN C J, WU Y, LI J W, et al. TBtools-Ⅱ: a “one for all, all for one” bioinformatics platform for biological big-data mining[J]. Molecular Plant, 2023, 16(11): 1733-1742. |
| [29] | CHEN S F, ZHOU Y Q, CHEN Y R, et al. Fastp: an ultra-fast all-in-one FASTQ preprocessor[J]. Bioinformatics, 2018, 34(17): i884-i890. |
| [30] | THAKUR V. RNA-seq data analysis for differential gene expression using HISAT2-stringtie-ballgown pipeline[J]. Methods in Molecular Biology, 2024, 2812: 101-113. |
| [31] | LIAO Y, SMYTH G K, SHI W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features[J]. Bioinformatics, 2014, 30(7): 923-930. |
| [32] | LIU D K, ZHANG C, ZHAO X, et al. Genome-wide analysis of the TCP gene family and their expression pattern in Cymbidium goeringii[J]. Frontiers in Plant Science, 2022, 13: 1068969. |
| [33] | SU C, ZHANG J, ZHENG C, et al. The orchid TCP gene family: genome-wide identification and expression analysis in different developmental stages and stress conditions[J]. BMC Plant Biology, 2019, 19(1): 1-16. |
| [34] | CAO S, ZHOU X, XIONG Z, et al. Genome-wide analysis of TCP transcription factor family in rye reveals their involvement in growth and abiotic stress responses[J]. International Journal of Molecular Sciences, 2021, 22(3): 1216. |
| [35] | LIN X, WU M, ZHANG Z, et al. Genome-wide analysis of TCP gene family in alfalfa (Medicago sativa L.) identifies roles in stem development and response to salt stress[J]. International Journal of Molecular Sciences, 2020, 21(6): 2021. |
| [36] | LIU H L, WU M, LI F, et al. TCP transcription factors in moso bamboo (Phyllostachys edulis): genome-wide identification and expression analysis[J]. Frontiers in Plant Science, 2018, 10(9):1263. |
| [37] | JEFFARES D C, PENKETT C J, BÄHLER J. Rapidly regulated genes are intron poor[J]. Trends in Genetics, 2008, 24(8): 375-378. |
| [38] | FLAGEL L E, WENDEL J F. Gene duplication and evolutionary novelty in plants[J]. New Phytologist, 2009, 183(3): 557-564. |
| [39] | Steven B. Cannon, Arvind Mitra, Andrew Baumgarten, et al. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana[J]. BMC Plant Biology, 2004, 4(1): 10. |
| [40] | LIU D H, LUO Y, HAN H, et al. Genome-wide analysis of citrus TCP transcription factors and their responses to abiotic stresses[J]. BMC Plant Biology, 2022, 22(1): 325. |
| [41] | XU Y, WANG Y, ZHANG L, et al. Genome-wide identification and characterization of the TCP transcription factor gene family in Broussonetia papyrifera[J]. Genes, 2021, 12(9): 1368. |
| [42] | VAN DE PEER Y, MAERE S, MEYER A. The evolutionary significance of ancient genome duplications[J]. Nature Reviews Genetics, 2009, 10(10): 725-732. |
| [43] | SHI G, XU Y, XIE K, et al. Evolution and expression analysis of TCP transcription factors in rice and Arabidopsis[J]. Journal of Plant Physiology, 2021, 266: 153523. |
| [44] | DHAKA N, BHARDWAJ V, SHARMA M K, et al. Evolving tale of TCPs: new paradigms and old lacunae[J]. Frontiers in Plant Science, 2017, 8: 479. |
| [45] | NATH U, CRAWFORD B C W, CARPENTER R, et al. Genetic control of surface curvature[J]. Science, 2003, 299(5611): 1404-1407. |
| [46] | DAVIÈRE J M, WILD M, REGNAULT T, et al. Class I TCP-DELLA interactions in inflorescence shoot apex determine plant height[J]. Current Biology, 2014, 24(16): 1923-1928. |
| [47] | 关紫微. 水稻耐逆基因OsTCP家族及OsSRZ1基因的功能研究[D]. 长沙: 湖南农业大学, 2021. |
| GUAN Z W. Functional study of OsTCP gene family and OsSRZ1 gene in rice stress tolerance[D]. Changsha: Hunan Agricultural University, 2021. | |
| [48] | JIANG Y, JIANG D, XIA M, et al. Genome-wide identification and expression analysis of the TCP gene family related to developmental and abiotic stress in ginger[J]. Plants, 2023, 12(19): 3389. |
| [49] | WU Q S, BAZZINI A A. Translation and mRNA stability control[J]. Annual Review of Biochemistry, 2023, 92: 227-245. |
| [50] | LEI N, YU X, LI S, et al. Phylogeny and expression pattern analysis of TCP transcription factors in cassava seedlings exposed to cold and/or drought stress[J]. Scientific Reports, 2017, 7: 10016. |
| [51] | YANG M F, HE G D, HOU Q D, et al. Systematic analysis and expression profiles of TCP gene family in Tartary buckwheat (Fagopyrum tataricum(L.) Gaertn.) revealed the potential function of FtTCP15 and FtTCP18 in response to abiotic stress[J]. BMC Genomics, 2022, 23(1): 415. |
| [52] | XU Y Z, LIU H L, GAO Y M, et al. The TCP transcription factor PeTCP10 modulates salt tolerance in transgenic Arabidopsis[J]. Plant Cell Reports, 2021, 40(10): 1971-1987. |
| [53] | SELAHATTIN D, AALT D J V D, ANDREA B, et al. Analysis of functional redundancies within the Arabidopsis TCP transcription factor family[J]. Journal of Experimental Botany, 2013, 64(18):5673-5685. |
| [1] | 胡心柔, 王梅, 张雅芬, 蔡为明, 金群力. 非生物胁迫对灵芝生长发育及其响应机制的影响[J]. 浙江农业学报, 2025, 37(5): 1182-1190. |
| [2] | 欧晋稳, 张古文, 冯志娟, 王斌, 卜远鹏, 徐钰, 茹磊, 刘娜, 龚亚明. 大豆海藻糖-6-磷酸磷酸酶基因GmTPP的鉴定及其在生长发育和非生物胁迫响应中的表达分析[J]. 浙江农业学报, 2024, 36(9): 2031-2041. |
| [3] | 张思懿, 崔博文, 王佳玲, 蔺吉祥, 杨青杰. 非生物胁迫下植物根系的生理与分子响应研究进展[J]. 浙江农业学报, 2024, 36(10): 2391-2401. |
| [4] | 张余, 金明伟, 任丽, 章毅颖, 赵洪, 刘昆, 邓姗, 褚云霞, 李寿国, 张靖立, 黄静艳, 陈海荣. 辣椒CaERF70的表达特征和转录自激活活性分析[J]. 浙江农业学报, 2024, 36(10): 2247-2256. |
| [5] | 寿伟松, 王铎, 沈佳, 许昕阳, 张跃建, 何艳军. 西瓜蔗糖转运蛋白SUT家族的鉴定及其在果实发育和逆境响应中的表达分析[J]. 浙江农业学报, 2024, 36(1): 94-102. |
| [6] | 熊雪, 赵丽娜, 杨森林, SAMIAH Arif, 张屹东. 甜瓜CmCIPK家族全基因组鉴定和逆境条件下的表达分析[J]. 浙江农业学报, 2021, 33(9): 1625-1639. |
| [7] | 何佳琦, 翟莹, 张军, 邱爽, 李铭杨, 赵艳, 张梅娟, 马天意. 大豆转录因子GmDof1.5的克隆与非生物胁迫诱导表达[J]. 浙江农业学报, 2021, 33(1): 1-7. |
| [8] | 邱文怡, 王诗雨, 李晓芳, 徐恒, 张华, 朱英, 王良超. MYB转录因子参与植物非生物胁迫响应与植物激素应答的研究进展[J]. 浙江农业学报, 2020, 32(7): 1317-1328. |
| [9] | 瓮巧云, 赵彦敏, 张贺, 宋晋辉, 马海莲, 袁进成, 刘颖慧. 玉米ZmREM基因的克隆与逆境胁迫后表达分析[J]. 浙江农业学报, 2016, 28(11): 1822-1827. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||