Acta Agriculturae Zhejiangensis ›› 2026, Vol. 38 ›› Issue (2): 269-283.DOI: 10.3969/j.issn.1004-1524.20241050
• Horticultural Science • Previous Articles Next Articles
Genome-wide analysis of the TCP gene family in Astragalus membranaceus (Fisch.) Bunge and its response to abiotic stress
MA Qiyanga( ), HUANG Xuelianb,*(
), HUANG Xuelianb,*( )
)
a. The First Clinical Medical College, Zhejiang Chinese Medical University Hangzhou 310053, China b. School of Basic Medical Sciences ,Zhejiang Chinese Medical University Hangzhou 310053, China
-
Received:2024-12-02Online:2026-02-25Published:2026-03-24
CLC Number:
Cite this article
MA Qiyang, HUANG Xuelian. Genome-wide analysis of the TCP gene family in Astragalus membranaceus (Fisch.) Bunge and its response to abiotic stress[J]. Acta Agriculturae Zhejiangensis, 2026, 38(2): 269-283.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.zjnyxb.cn/EN/10.3969/j.issn.1004-1524.20241050
| 基因 Gene | 正向引物(5'→3') Forward primer (5'→3') | 反向引物(5'→3') Reverse primer (5'→3') |
|---|---|---|
| Actin | CTCTCTCAGCACCTTCGAGCAG | TCCACATACAACCGCTCCACTG |
| AmTCP8 | CCAGGCGGAACCGAGCATTATTG | TCGCTTCCTGCTGCTGATGATGA |
| AmTCP16 | AACGGCGATGATGGCGATGATG | TGCTGCTGCTGCTGCTGATG |
| AmTCP18 | GCGGCAGCGGCAACTATAACAA | GCTCGGAATATGATGATGATGCGGAAT |
| AmTCP19 | AGCAGCAGCAGCAGCAGAAC | CGTTGTTGCCGGTATCCAGGTT |
| AmTCP23 | AGCCGCACCAACTGGAACATTAC | TCTGATGCTGATGCTGGTTCTGATG |
Table 1 Primers for RT-qPCR
| 基因 Gene | 正向引物(5'→3') Forward primer (5'→3') | 反向引物(5'→3') Reverse primer (5'→3') |
|---|---|---|
| Actin | CTCTCTCAGCACCTTCGAGCAG | TCCACATACAACCGCTCCACTG |
| AmTCP8 | CCAGGCGGAACCGAGCATTATTG | TCGCTTCCTGCTGCTGATGATGA |
| AmTCP16 | AACGGCGATGATGGCGATGATG | TGCTGCTGCTGCTGCTGATG |
| AmTCP18 | GCGGCAGCGGCAACTATAACAA | GCTCGGAATATGATGATGATGCGGAAT |
| AmTCP19 | AGCAGCAGCAGCAGCAGAAC | CGTTGTTGCCGGTATCCAGGTT |
| AmTCP23 | AGCCGCACCAACTGGAACATTAC | TCTGATGCTGATGCTGGTTCTGATG |
| 蛋白名 Protein name | 基因名 Gene name | 氨基酸数量 Amino acid quantity | 相对分子质量/ku Relative molecular weight/ku | 等电点 Isoelectric point | 不稳定系数 Instability index | 脂肪指数 Aliphatic index | 亲水性 Hydrophilicity |
|---|---|---|---|---|---|---|---|
| AmTCP1 | Amem01G08650.1 | 334 | 36.25 | 9.62 | 61.72 | 72.78 | -0.355 |
| AmTCP2 | Amem01G26990.1 | 353 | 37.44 | 5.99 | 52.15 | 67.73 | -0.460 |
| AmTCP3 | Amem01G29700.1 | 412 | 44.15 | 7.97 | 67.32 | 58.33 | -0.720 |
| AmTCP4 | Amem01G30870.1 | 349 | 39.08 | 8.44 | 53.65 | 55.62 | -0.756 |
| AmTCP5 | Amem01G31480.1 | 342 | 39.30 | 9.16 | 57.79 | 55.85 | -1.099 |
| AmTCP6 | Amem01G39660.1 | 329 | 35.68 | 6.21 | 43.87 | 57.63 | -0.681 |
| AmTCP7 | Amem02G01750.1 | 261 | 27.81 | 9.56 | 48.82 | 70.73 | -0.475 |
| AmTCP8 | Amem02G07350.1 | 233 | 25.13 | 9.10 | 52.77 | 70.52 | -0.566 |
| AmTCP9 | Amem02G13390.1 | 203 | 21.44 | 8.44 | 61.98 | 68.82 | -0.265 |
| AmTCP10 | Amem02G30910.1 | 344 | 36.12 | 5.46 | 54.31 | 65.00 | -0.358 |
| AmTCP11 | Amem02G31020.1 | 344 | 36.13 | 5.46 | 53.03 | 65.00 | -0.358 |
| AmTCP12 | Amem03G33990.1 | 375 | 41.03 | 6.90 | 66.54 | 52.88 | -0.784 |
| AmTCP13 | Amem03G35070.1 | 430 | 47.97 | 7.42 | 47.37 | 59.35 | -0.770 |
| AmTCP14 | Amem04G15690.1 | 417 | 44.13 | 6.90 | 56.30 | 57.15 | -0.661 |
| AmTCP15 | Amem05G34130.3 | 450 | 49.85 | 7.21 | 52.74 | 52.27 | -0.986 |
| AmTCP16 | Amem06G00490.1 | 494 | 52.19 | 6.15 | 62.09 | 50.63 | -0.856 |
| AmTCP17 | Amem06G06270.2 | 355 | 39.83 | 8.56 | 58.10 | 55.55 | -1.028 |
| AmTCP18 | Amem06G19960.1 | 177 | 20.64 | 10.50 | 49.17 | 69.44 | -0.921 |
| AmTCP19 | Amem06G25360.1 | 348 | 37.92 | 5.67 | 57.14 | 59.17 | -0.719 |
| AmTCP20 | Amem07G03630.1 | 358 | 40.03 | 8.28 | 49.00 | 71.84 | -0.665 |
| AmTCP21 | Amem07G04880.1 | 391 | 44.30 | 9.12 | 51.58 | 66.14 | -0.874 |
| AmTCP22 | Amem09G02360.1 | 358 | 40.20 | 8.26 | 54.21 | 55.89 | -0.885 |
| AmTCP23 | Amem09G03560.1 | 322 | 35.47 | 9.62 | 52.23 | 58.79 | -0.730 |
| AmTCP24 | Amem09G20040.1 | 362 | 40.59 | 8.87 | 44.80 | 69.53 | -0.735 |
Table 2 Physicochemical characteristics of 24 AmTCP genes and their encoded proteins
| 蛋白名 Protein name | 基因名 Gene name | 氨基酸数量 Amino acid quantity | 相对分子质量/ku Relative molecular weight/ku | 等电点 Isoelectric point | 不稳定系数 Instability index | 脂肪指数 Aliphatic index | 亲水性 Hydrophilicity |
|---|---|---|---|---|---|---|---|
| AmTCP1 | Amem01G08650.1 | 334 | 36.25 | 9.62 | 61.72 | 72.78 | -0.355 |
| AmTCP2 | Amem01G26990.1 | 353 | 37.44 | 5.99 | 52.15 | 67.73 | -0.460 |
| AmTCP3 | Amem01G29700.1 | 412 | 44.15 | 7.97 | 67.32 | 58.33 | -0.720 |
| AmTCP4 | Amem01G30870.1 | 349 | 39.08 | 8.44 | 53.65 | 55.62 | -0.756 |
| AmTCP5 | Amem01G31480.1 | 342 | 39.30 | 9.16 | 57.79 | 55.85 | -1.099 |
| AmTCP6 | Amem01G39660.1 | 329 | 35.68 | 6.21 | 43.87 | 57.63 | -0.681 |
| AmTCP7 | Amem02G01750.1 | 261 | 27.81 | 9.56 | 48.82 | 70.73 | -0.475 |
| AmTCP8 | Amem02G07350.1 | 233 | 25.13 | 9.10 | 52.77 | 70.52 | -0.566 |
| AmTCP9 | Amem02G13390.1 | 203 | 21.44 | 8.44 | 61.98 | 68.82 | -0.265 |
| AmTCP10 | Amem02G30910.1 | 344 | 36.12 | 5.46 | 54.31 | 65.00 | -0.358 |
| AmTCP11 | Amem02G31020.1 | 344 | 36.13 | 5.46 | 53.03 | 65.00 | -0.358 |
| AmTCP12 | Amem03G33990.1 | 375 | 41.03 | 6.90 | 66.54 | 52.88 | -0.784 |
| AmTCP13 | Amem03G35070.1 | 430 | 47.97 | 7.42 | 47.37 | 59.35 | -0.770 |
| AmTCP14 | Amem04G15690.1 | 417 | 44.13 | 6.90 | 56.30 | 57.15 | -0.661 |
| AmTCP15 | Amem05G34130.3 | 450 | 49.85 | 7.21 | 52.74 | 52.27 | -0.986 |
| AmTCP16 | Amem06G00490.1 | 494 | 52.19 | 6.15 | 62.09 | 50.63 | -0.856 |
| AmTCP17 | Amem06G06270.2 | 355 | 39.83 | 8.56 | 58.10 | 55.55 | -1.028 |
| AmTCP18 | Amem06G19960.1 | 177 | 20.64 | 10.50 | 49.17 | 69.44 | -0.921 |
| AmTCP19 | Amem06G25360.1 | 348 | 37.92 | 5.67 | 57.14 | 59.17 | -0.719 |
| AmTCP20 | Amem07G03630.1 | 358 | 40.03 | 8.28 | 49.00 | 71.84 | -0.665 |
| AmTCP21 | Amem07G04880.1 | 391 | 44.30 | 9.12 | 51.58 | 66.14 | -0.874 |
| AmTCP22 | Amem09G02360.1 | 358 | 40.20 | 8.26 | 54.21 | 55.89 | -0.885 |
| AmTCP23 | Amem09G03560.1 | 322 | 35.47 | 9.62 | 52.23 | 58.79 | -0.730 |
| AmTCP24 | Amem09G20040.1 | 362 | 40.59 | 8.87 | 44.80 | 69.53 | -0.735 |
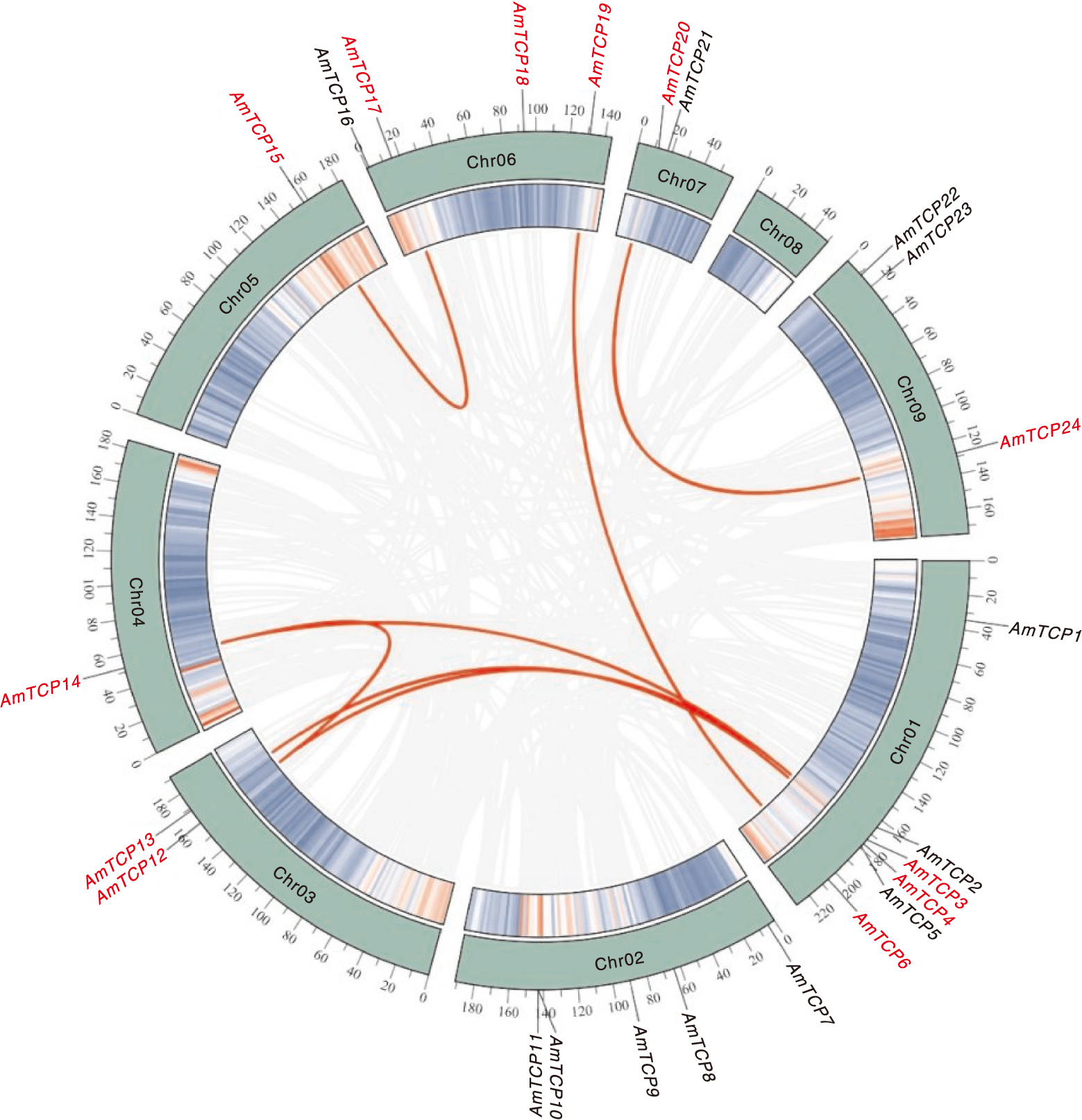
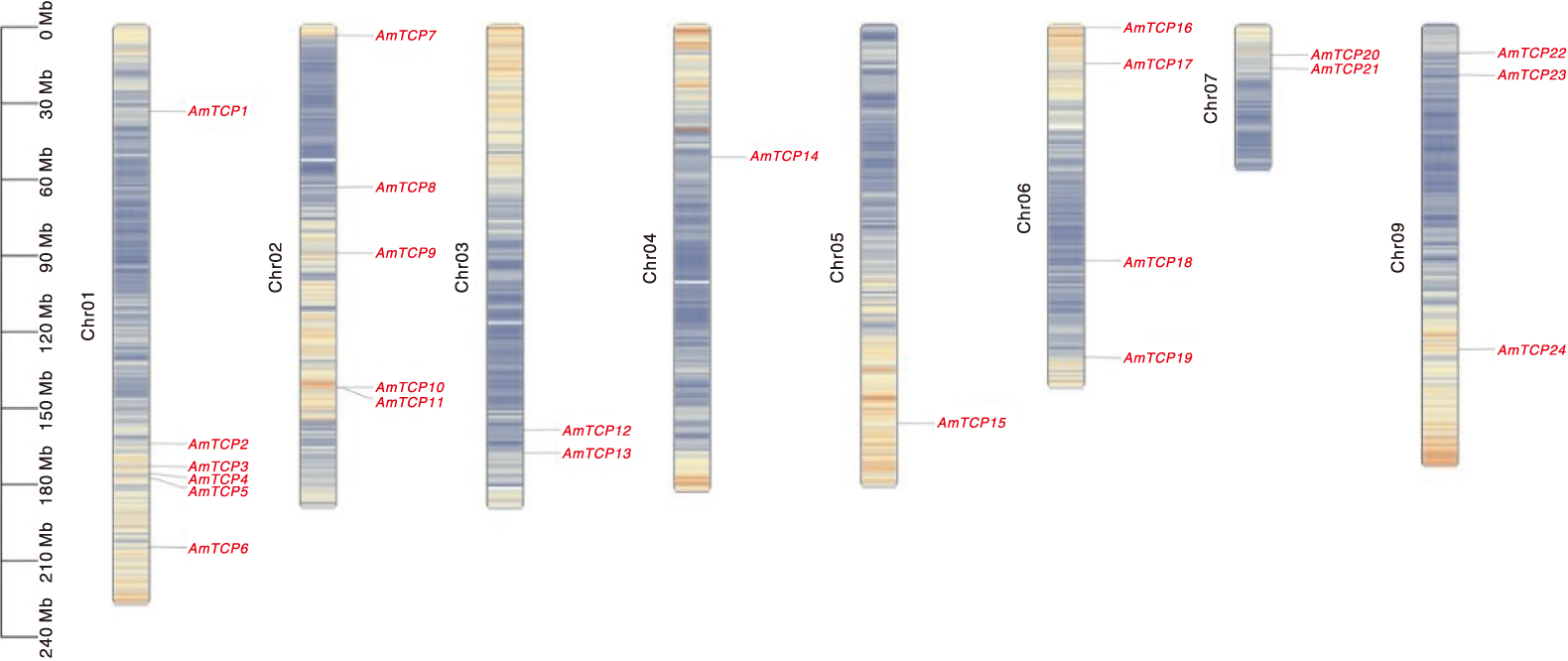
Fig.5 Colinearity of the AmTCP gene The green line in the inner ring represents the gene density per chromosome, co-lined genes are indicated by light gray lines, and intraspecific co-lined gene pairs of AmTCP are labeled with red curves and the font is also labeled in red.
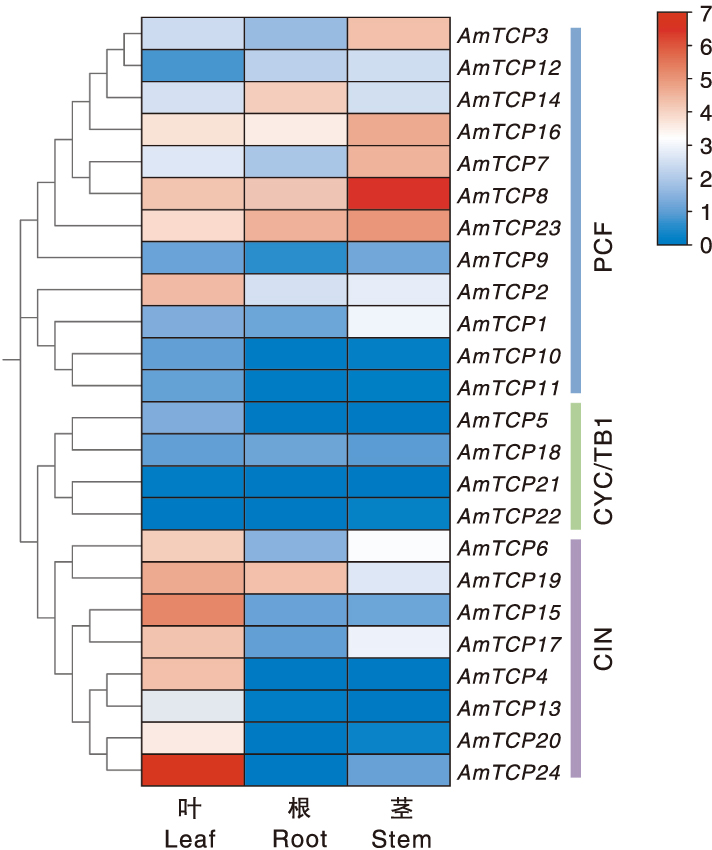
Fig.8 AmTCP gene expression in different tissues Color bars represent normalised expression levels (log2 converted fold change). Red and blue indicate up- and down-regulated genes, and white indicates no expression. The blue, green and purple bands on the right represent the PCF, CYC/TB1 and CIN subfamilies.
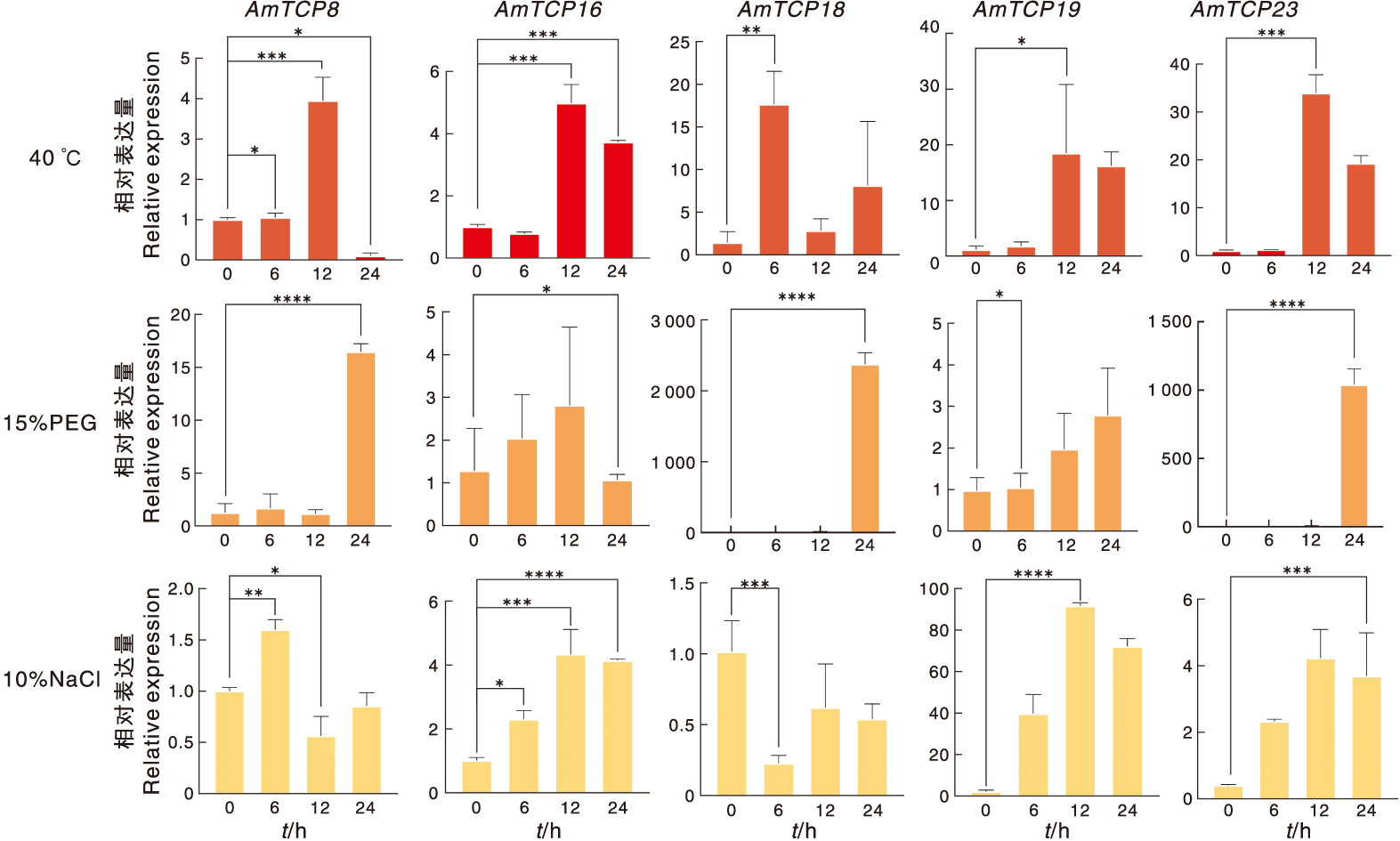
Fig.9 Expression of AmTCP genes under abiotic stress High temperature (40 ℃), drought (15% PEG), salt (10% NaCl). “*” indicates a significant difference at the 0.05 level between different treatments, “**” indicates a significant difference at the 0.01 level between different treatments, “***” indicates a significant difference at the 0.000 5 level between different treatments, “****” indicates a significant difference at the 0.000 1 level between different treatments.
| [1] | CUBAS P, LAUTER N, DOEBLEY J, et al. The TCP domain: a motif found in proteins regulating plant growth and development[J]. The Plant Journal, 1999, 18(2): 215-222. |
| [2] | DOEBLEY J, STEC A, HUBBARD L. The evolution of apical dominance in maize[J]. Nature, 1997, 386(6624): 485-488. |
| [3] | LUO D, CARPENTER R, VINCENT C, et al. Origin of floral asymmetry in Antirrhinum[J]. Nature, 1996, 383(6603): 794-799. |
| [4] | KOSUGI S, OHASHI Y. PCF1 and PCF2 specifically bind to cis elements in the rice proliferating cell nuclear antigen gene[J]. The Plant Cell, 1997, 9(9): 1607-1619. |
| [5] | AGGARWAL P, DAS GUPTA M, JOSEPH A P, et al. Identification of specific DNA binding residues in the TCP family of transcription factors in Arabidopsis[J]. The Plant Cell, 2010, 22(4): 1174-1189. |
| [6] | MARTÍN-TRILLO M, CUBAS P. TCP genes: a family snapshot ten years later[J]. Trends in Plant Science, 2010, 15(1): 31-39. |
| [7] | PARAPUNOVA V, BUSSCHER M, BUSSCHER-LANGE J, et al. Identification, cloning and characterization of the tomato TCP transcription factor family[J]. BMC Plant Biology, 2014, 14(1): 157. |
| [8] | LI Z, LI B, SHEN W, et al. Comprehensive genomic identification and expression analysis of TCP transcription factors in kiwifruit (Actinidia chinensis)[J]. BMC Genomics, 2019, 19(1): 271. |
| [9] | QI X H, SUN S L, XU P, et al. Genome-wide identification and expression analysis of the TCP transcription factor genes in Capsicum annuum L.[J]. International Journal of Molecular Sciences, 2016, 17(6): 702. |
| [10] | WANG Y, CHEN D, WANG D, et al. Genome-wide analysis of TCP transcription factor gene family in Medicago sativa reveals significant roles in shoot lateral organ development[J]. BMC Plant Biology, 2015, 15: 29. |
| [11] | LIU C, LYU T T, SHEN Y H, et al. Genome-wide identification and integrated analysis of TCP genes controlling ginsenoside biosynthesis in Panax ginseng[J]. BMC Plant Biology, 2024, 24(1): 47. |
| [12] | KOYAMA T, SATO F, OHME-TAKAGI M. Roles of miR319 and TCP transcription factors in leaf development[J]. Plant Physiology, 2017, 175(2): 874-885. |
| [13] | GUAN P, RIPOLL J J, WANG R, et al. Interacting TCP and NLP transcription factors control plant responses to nitrate availability[J]. Proceedings of the National Academy of Sciences of the United States of America, 2017, 114(9): 2419-2424. |
| [14] | MUKHOPADHYAY P, TYAGI A K. OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways[J]. Scientific Reports, 2015, 5: 9998. |
| [15] | SHARMA R, SINGH G, BHATTACHARYA S, et al. A genome-wide survey of TCP transcription factors provides insights into their functional roles in chickpea (Cicer arietinum L.)[J]. Scientific Reports, 2018, 8(1): 8455. |
| [16] | WU X Y, LI J Z, WEN X H, et al. Genome-wide identification of the TCP gene family in Chrysanthemum lavandulifolium and its homologs expression patterns during flower development in different Chrysanthemum species[J]. Frontiers in Plant Science, 2023, 14(1): 1276123. |
| [17] | CHAI W, JIANG P, HUANG G, et al. ZmTCP42, a TCP transcription factor from maize, confers drought tolerance in transgenic Arabidopsis[J]. International Journal of Molecular Sciences, 2021, 22(4): 2024. |
| [18] | MA J, WANG Q L, SUN R R, et al. Genome-wide identification and expression analysis of TCP transcription factors in Gossypium raimondii[J]. Scientific Reports, 2014, 4: 6645. |
| [19] | 后叶虎, 邱璐, 靳晓杰, 等. 基于化学生物信息学方法探讨黄芪通过调节能量代谢发挥“补气”功效的物质基础和分子机制[J]. 中国现代应用药学, 2023, 40(14): 1906. |
| HOU Y H, QIU L, JIN X J, et al. Exploration on material basis and molecular mechanism of astragali radix exerting the effect of “Qi-invigorating” through regulating energy metabolism based on chemo-bio informatics methods[J]. Chinese Journal of Modern Applied Pharmacy, 2023, 40(14): 1906. | |
| [20] | 余意, 胡明华, 张丹丹, 等. 黄芪多糖对气虚大鼠的补气作用及其机制探讨[J]. 中药新药与临床药理, 2021, 32(4): 505. |
| YU Y, HU M H, ZHANG D D, et al. The “Qi-invigorating” effect and possible underlying mechanism of astragalus polysaccharides on rats with Qi-deficiency[J]. Chinese Journal of New Drugs and Clinical Remedies, 2021, 32(4): 505. | |
| [21] | 周文静, 柴智, 刘佳欣, 等. 黄芪补气功用探微[J]. 中国民间疗法, 2021, 29(20): 1. |
| ZHOU W J, CHAI Z, LIU J X, et al. Study on the function of “Qi-invigorating” with Astragalus membranaceus[J]. Chinese Folk Therapy, 2021, 29(20): 1. | |
| [22] | 张倩倩, 李光跃, 苏优拉, 等. 干旱胁迫对蒙古黄芪和膜荚黄芪不同器官黄酮类成分积累的影响[J]. 西北植物学报, 2020, 40(7): 1201. |
| ZHANG Q Q, LI G Y, SU Y L, et al. Effects of drought stress on flavonoid accumulation in different organs of Astragalus mongholicus and Astragalus membranaceus[J]. Acta Botanica Boreali-Occidentalia Sinica, 2020, 40(7): 1201. | |
| [23] | 王楠, 高静, 岳正刚, 等. 黄芪种子萌发对光温条件和旱盐胁迫的响应特征[J]. 草业学报, 2018, 27(1): 106. |
| WANG N, GAO J, YUE Z G, et al. Response characteristics of Astragalus seed germination to light-temperature conditions and drought-salt stress[J]. Acta Prataculturae Sinica, 2018, 27(1): 106. | |
| [24] | 刘洋. 应用代谢组学对比研究两种黄芪响应UV-B和干旱胁迫的代谢基础[D]. 哈尔滨: 东北林业大学, 2016. |
| LIU Y. Comparative study on the metabolic basis of two Astragalus membranaceus species in response to UV-B and drought stress based on metabolomics[D]. Harbin: Northeast Forestry University, 2016. | |
| [25] | WANG J L, WANG H W, CAO Y N, et al. Comprehensive evolutionary analysis of the TCP gene family: Further insights for its origin, expansion, and diversification[J]. Frontiers in Plant Science, 2022, 13: 994567. |
| [26] | ZHOU H Y, HWARARI D, MA H Y, et al. Genomic survey of TCP transcription factors in plants: phylogenomics, evolution and their biology[J]. Frontiers in Genetics, 2022, 13: 1060546. |
| [27] | FAN H J, CHAI Z, YANG X K, et al. Chromosome-scale genome assembly of Astragalus membranaceus using PacBio and Hi-C technologies[J]. Scientific Data, 2024, 11(1):1071. |
| [28] | CHEN C J, WU Y, LI J W, et al. TBtools-Ⅱ: a “one for all, all for one” bioinformatics platform for biological big-data mining[J]. Molecular Plant, 2023, 16(11): 1733-1742. |
| [29] | CHEN S F, ZHOU Y Q, CHEN Y R, et al. Fastp: an ultra-fast all-in-one FASTQ preprocessor[J]. Bioinformatics, 2018, 34(17): i884-i890. |
| [30] | THAKUR V. RNA-seq data analysis for differential gene expression using HISAT2-stringtie-ballgown pipeline[J]. Methods in Molecular Biology, 2024, 2812: 101-113. |
| [31] | LIAO Y, SMYTH G K, SHI W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features[J]. Bioinformatics, 2014, 30(7): 923-930. |
| [32] | LIU D K, ZHANG C, ZHAO X, et al. Genome-wide analysis of the TCP gene family and their expression pattern in Cymbidium goeringii[J]. Frontiers in Plant Science, 2022, 13: 1068969. |
| [33] | SU C, ZHANG J, ZHENG C, et al. The orchid TCP gene family: genome-wide identification and expression analysis in different developmental stages and stress conditions[J]. BMC Plant Biology, 2019, 19(1): 1-16. |
| [34] | CAO S, ZHOU X, XIONG Z, et al. Genome-wide analysis of TCP transcription factor family in rye reveals their involvement in growth and abiotic stress responses[J]. International Journal of Molecular Sciences, 2021, 22(3): 1216. |
| [35] | LIN X, WU M, ZHANG Z, et al. Genome-wide analysis of TCP gene family in alfalfa (Medicago sativa L.) identifies roles in stem development and response to salt stress[J]. International Journal of Molecular Sciences, 2020, 21(6): 2021. |
| [36] | LIU H L, WU M, LI F, et al. TCP transcription factors in moso bamboo (Phyllostachys edulis): genome-wide identification and expression analysis[J]. Frontiers in Plant Science, 2018, 10(9):1263. |
| [37] | JEFFARES D C, PENKETT C J, BÄHLER J. Rapidly regulated genes are intron poor[J]. Trends in Genetics, 2008, 24(8): 375-378. |
| [38] | FLAGEL L E, WENDEL J F. Gene duplication and evolutionary novelty in plants[J]. New Phytologist, 2009, 183(3): 557-564. |
| [39] | Steven B. Cannon, Arvind Mitra, Andrew Baumgarten, et al. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana[J]. BMC Plant Biology, 2004, 4(1): 10. |
| [40] | LIU D H, LUO Y, HAN H, et al. Genome-wide analysis of citrus TCP transcription factors and their responses to abiotic stresses[J]. BMC Plant Biology, 2022, 22(1): 325. |
| [41] | XU Y, WANG Y, ZHANG L, et al. Genome-wide identification and characterization of the TCP transcription factor gene family in Broussonetia papyrifera[J]. Genes, 2021, 12(9): 1368. |
| [42] | VAN DE PEER Y, MAERE S, MEYER A. The evolutionary significance of ancient genome duplications[J]. Nature Reviews Genetics, 2009, 10(10): 725-732. |
| [43] | SHI G, XU Y, XIE K, et al. Evolution and expression analysis of TCP transcription factors in rice and Arabidopsis[J]. Journal of Plant Physiology, 2021, 266: 153523. |
| [44] | DHAKA N, BHARDWAJ V, SHARMA M K, et al. Evolving tale of TCPs: new paradigms and old lacunae[J]. Frontiers in Plant Science, 2017, 8: 479. |
| [45] | NATH U, CRAWFORD B C W, CARPENTER R, et al. Genetic control of surface curvature[J]. Science, 2003, 299(5611): 1404-1407. |
| [46] | DAVIÈRE J M, WILD M, REGNAULT T, et al. Class I TCP-DELLA interactions in inflorescence shoot apex determine plant height[J]. Current Biology, 2014, 24(16): 1923-1928. |
| [47] | 关紫微. 水稻耐逆基因OsTCP家族及OsSRZ1基因的功能研究[D]. 长沙: 湖南农业大学, 2021. |
| GUAN Z W. Functional study of OsTCP gene family and OsSRZ1 gene in rice stress tolerance[D]. Changsha: Hunan Agricultural University, 2021. | |
| [48] | JIANG Y, JIANG D, XIA M, et al. Genome-wide identification and expression analysis of the TCP gene family related to developmental and abiotic stress in ginger[J]. Plants, 2023, 12(19): 3389. |
| [49] | WU Q S, BAZZINI A A. Translation and mRNA stability control[J]. Annual Review of Biochemistry, 2023, 92: 227-245. |
| [50] | LEI N, YU X, LI S, et al. Phylogeny and expression pattern analysis of TCP transcription factors in cassava seedlings exposed to cold and/or drought stress[J]. Scientific Reports, 2017, 7: 10016. |
| [51] | YANG M F, HE G D, HOU Q D, et al. Systematic analysis and expression profiles of TCP gene family in Tartary buckwheat (Fagopyrum tataricum(L.) Gaertn.) revealed the potential function of FtTCP15 and FtTCP18 in response to abiotic stress[J]. BMC Genomics, 2022, 23(1): 415. |
| [52] | XU Y Z, LIU H L, GAO Y M, et al. The TCP transcription factor PeTCP10 modulates salt tolerance in transgenic Arabidopsis[J]. Plant Cell Reports, 2021, 40(10): 1971-1987. |
| [53] | SELAHATTIN D, AALT D J V D, ANDREA B, et al. Analysis of functional redundancies within the Arabidopsis TCP transcription factor family[J]. Journal of Experimental Botany, 2013, 64(18):5673-5685. |
| [1] | HU Xinrou, WANG Mei, ZHANG Yafen, CAI Weiming, JIN Qunli. Effect of abiotic stress on growth development and response mechanism of Ganoderma [J]. Acta Agriculturae Zhejiangensis, 2025, 37(5): 1182-1190. |
| [2] | OU Jinwen, ZHANG Guwen, FENG Zhijuan, WANG Bin, BU Yuanpeng, XU Yu, RU Lei, LIU Na, GONG Yaming. Identification of soybean trehalose-6-phosphate phosphatase gene GmTPP and its expression analysis in growth and abiotic stress response [J]. Acta Agriculturae Zhejiangensis, 2024, 36(9): 2031-2041. |
| [3] | ZHANG Siyi, CUI Bowen, WANG Jialing, LIN Jixiang, YANG Qingjie. Research progress on physiological and molecular responses of plant roots under abiotic stress [J]. Acta Agriculturae Zhejiangensis, 2024, 36(10): 2391-2401. |
| [4] | ZHANG Yu, JIN Mingwei, REN Li, ZHANG Yiying, ZHAO Hong, LIU Kun, DENG Shan, CHU Yunxia, LI Shouguo, ZHANG Jingli, HUANG Jingyan, CHEN Hairong. Expression patterns and transcriptional autoactivation analysis of CaERF70 in chili pepper [J]. Acta Agriculturae Zhejiangensis, 2024, 36(10): 2247-2256. |
| [5] | SHOU Weisong, WANG Duo, SHEN Jia, XU Xinyang, ZHANG Yuejian, HE Yanjun. Identification and expression analysis of sucrose transporter SUT family in watermelon in fruit development and stress responses [J]. Acta Agriculturae Zhejiangensis, 2024, 36(1): 94-102. |
| [6] | JIA Liqiang, ZHAO Qiufang, CHEN Shu, DING Bo. Expression analysis of bZIP G subfamily genes in maize [J]. Acta Agriculturae Zhejiangensis, 2022, 34(2): 221-231. |
| [7] | XIONG Xue, ZHAO Lina, YANG Senlin, SAMIAH Arif, ZHANG Yidong. Genome-wide identification of CmCIPK family and its expression analysis under abiotic stress in melon [J]. Acta Agriculturae Zhejiangensis, 2021, 33(9): 1625-1639. |
| [8] | HE Jiaqi, ZHAI Ying, ZHANG Jun, QIU Shuang, LI Mingyang, ZHAO Yan, ZHANG Meijuan, MA Tianyi. Cloning and expression analysis of GmDof1.5 in soybean under abiotic stress [J]. Acta Agriculturae Zhejiangensis, 2021, 33(1): 1-7. |
| [9] | QIU Wenyi, WANG Shiyu, LI Xiaofang, XU Heng, ZHANG Hua, ZHU Ying, WANG Liangchao. Functions of plant MYB transcription factors in response to abiotic stress and plant hormones [J]. , 2020, 32(7): 1317-1328. |
| [10] | JIANG Yaoyao, LI Jing, CAI Nianjun, CHEN Jianping, ZHANG Hengmu. Cloning and expression analysis of a fibrillin gene in plant [J]. , 2019, 31(9): 1399-1404. |
| [11] | LIU Huijie, XU Heng, QIU Wenyi, LI Xiaofang, ZHANG Hua, ZHU Ying, LI Chunshou, WANG Liangchao. Roles of bZIP transcription factors in plant growth and development and abiotic stress response [J]. , 2019, 31(7): 1205-1214. |
| [12] | WENG Qiao-yun, ZHAO Yan-min, ZHANG He, SONG Jin-hui, MA Hai-lian, YUAN Jin-cheng, LIU Ying-hui. Cloning of ZmREM and its expression analysis in maize under stress [J]. , 2016, 28(11): 1822-1827. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||