Acta Agriculturae Zhejiangensis ›› 2026, Vol. 38 ›› Issue (1): 1-16.DOI: 10.3969/j.issn.1004-1524.20250086
• Crop Science • Previous Articles Next Articles
Identification of common genes for salt and cadmium tolerance in soybean
XU Yan1,2( ), LI Sujuan3, CHEN Guang2, XU Shengchun1,2,4, WANG Jian2,*(
), LI Sujuan3, CHEN Guang2, XU Shengchun1,2,4, WANG Jian2,*( )
)
- 1. College of Advanced Agricultural Science, Zhejiang A&F University, Hangzhou 311300, China
2. Institute of Digital Agriculture, Zhejiang Academy of Agricultural Sciences, Hangzhou 310021, China
3. State Key Laboratory for Quality and Safety of Agro-Products, Zhejiang Academy of Agricultural Sciences, Hangzhou 310021, China
4. Xianghu Laboratory, Hangzhou 311258, China
-
Received:2025-02-07Online:2026-01-25Published:2026-02-11
CLC Number:
Cite this article
XU Yan, LI Sujuan, CHEN Guang, XU Shengchun, WANG Jian. Identification of common genes for salt and cadmium tolerance in soybean[J]. Acta Agriculturae Zhejiangensis, 2026, 38(1): 1-16.
share this article
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.zjnyxb.cn/EN/10.3969/j.issn.1004-1524.20250086
| 基因Gene | 正向引物序列(5'→3') Forward primer sequence(5'→3') | 反向引物序列(5'→3') Reverse primer sequence(5'→3') |
|---|---|---|
| GmEF1b | CCACTGCTGAAGAAGATGATGATG | AAGGACAGAAGACTTGCCACTC |
| GmDehydrin | AGGAAGGAACATCGTCAGCA | TGACAAGACACTGTACGTACG |
| GmSSP | CCACCTCAGGAGTCTCAGAA | CCCGCAAAAGTTTCGTGACT |
| GmGF14 | ACGTTGGGAGAGGAATCATACA | GCATTCAACACCTTCTCCCT |
| GmPAP85 | AGCAGAAAGAGGAGGGGAAC | AGCAGACAGTTGAAGTACACA |
| GmHUP54 | ATGCCTAGGATTGACAGCGA | AGCAGAGTCAGCACCATCAT |
| GmMET2 | TCGAGAGTGCTGAAATGGGT | ACACACCCATCACAAGTCCA |
Table 1 Primers for qRT-PCR analysis of candidate genes associated with salt- and cadmium-tolerance
| 基因Gene | 正向引物序列(5'→3') Forward primer sequence(5'→3') | 反向引物序列(5'→3') Reverse primer sequence(5'→3') |
|---|---|---|
| GmEF1b | CCACTGCTGAAGAAGATGATGATG | AAGGACAGAAGACTTGCCACTC |
| GmDehydrin | AGGAAGGAACATCGTCAGCA | TGACAAGACACTGTACGTACG |
| GmSSP | CCACCTCAGGAGTCTCAGAA | CCCGCAAAAGTTTCGTGACT |
| GmGF14 | ACGTTGGGAGAGGAATCATACA | GCATTCAACACCTTCTCCCT |
| GmPAP85 | AGCAGAAAGAGGAGGGGAAC | AGCAGACAGTTGAAGTACACA |
| GmHUP54 | ATGCCTAGGATTGACAGCGA | AGCAGAGTCAGCACCATCAT |
| GmMET2 | TCGAGAGTGCTGAAATGGGT | ACACACCCATCACAAGTCCA |
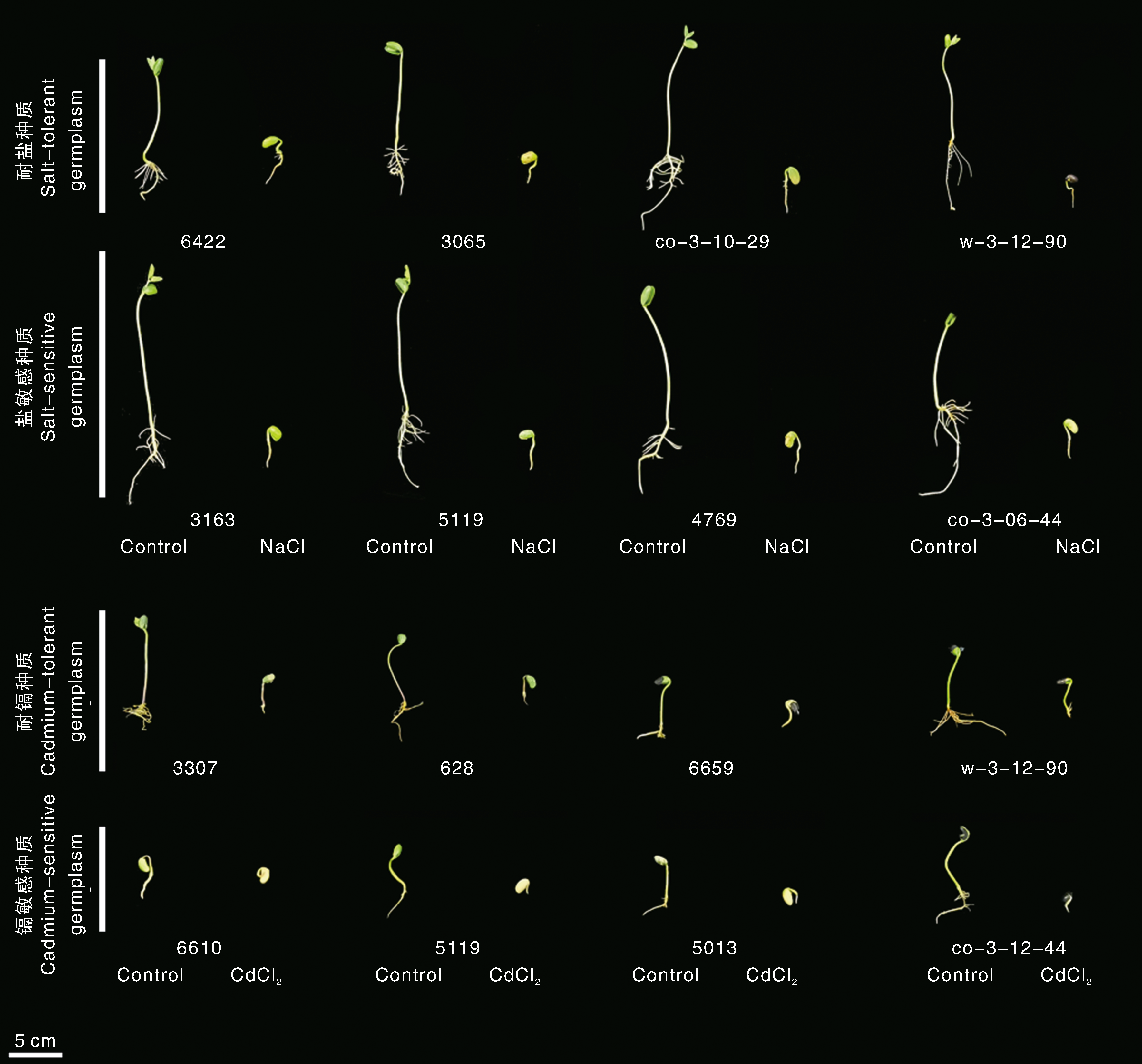
Fig.1 Physiological performance of different soybean seedlings under salt or cadmium stress Physiological responses of soybean seedlings of salt/Cd-tolerant and salt/Cd-sensitive varieties under salt/Cd stress for 8 days. NaCl, Salt stress; Cd, Cadmium stress; Control, Normal growth.
| 处理 Treatment | PL/cm | RL/cm | RL/PL | SW/g | RW/g | GR | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | |
| 正常条件 CK | 10.26± 0.31 a | 24.91 | 7.52± 0.16 a | 18.27 | 0.81± 0.02 b | 23.83 | 0.76± 0.02 a | 24.14 | 0.24± 0.01 a | 31.58 | 0.97± 0.01 a | 6.15 |
| 盐胁迫 Salt stress | 1.03± 0.02 b | 16.75 | 1.86± 0.05 b | 23.06 | 1.92± 0.05 a | 24.63 | 0.41± 0.01 b | 31.97 | 0.05± 0.00 b | 46.62 | 0.92± 0.01 b | 10.56 |
| 镉胁迫 Cd stress | 0.88± 0.03 b | 37.75 | 0.30± 0.04 c | 133.61 | 0.30± 0.04 c | 134.28 | 0.41± 0.01 b | 30.95 | 0.03± 0.01 b | 353.39 | 0.75± 0.02 c | 31.25 |
Table 2 Physiological performance of 50 soybean germplasms during seedling stage under normal conditions and salt/cadmium stress
| 处理 Treatment | PL/cm | RL/cm | RL/PL | SW/g | RW/g | GR | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | 均值 Mean | CV/% | |
| 正常条件 CK | 10.26± 0.31 a | 24.91 | 7.52± 0.16 a | 18.27 | 0.81± 0.02 b | 23.83 | 0.76± 0.02 a | 24.14 | 0.24± 0.01 a | 31.58 | 0.97± 0.01 a | 6.15 |
| 盐胁迫 Salt stress | 1.03± 0.02 b | 16.75 | 1.86± 0.05 b | 23.06 | 1.92± 0.05 a | 24.63 | 0.41± 0.01 b | 31.97 | 0.05± 0.00 b | 46.62 | 0.92± 0.01 b | 10.56 |
| 镉胁迫 Cd stress | 0.88± 0.03 b | 37.75 | 0.30± 0.04 c | 133.61 | 0.30± 0.04 c | 134.28 | 0.41± 0.01 b | 30.95 | 0.03± 0.01 b | 353.39 | 0.75± 0.02 c | 31.25 |
| 性状 Trait | 主成分1 PC1 | 主成分2 PC2 | 主成分3 PC3 |
|---|---|---|---|
| PL | 0.86 | -0.12 | 0.08 |
| RL | 0.35 | 0.82 | -0.12 |
| GR | -0.49 | 0.84 | -0.09 |
| RL/PL | 0.42 | 0.67 | 0.24 |
| RW | 0.84 | -0.04 | 0.20 |
| SW | -0.37 | 0.04 | 0.90 |
| 特征值Eigenvalue | 2.09 | 1.81 | 0.93 |
| 贡献率/%Contribution rate/% | 34.85 | 30.12 | 15.52 |
| 累计贡献率/% | 34.85 | 64.97 | 80.49 |
| Cumulative contribution rate/% |
Table 3 Contribution of variables to principal components under salt stress across 50 soybean germplasms
| 性状 Trait | 主成分1 PC1 | 主成分2 PC2 | 主成分3 PC3 |
|---|---|---|---|
| PL | 0.86 | -0.12 | 0.08 |
| RL | 0.35 | 0.82 | -0.12 |
| GR | -0.49 | 0.84 | -0.09 |
| RL/PL | 0.42 | 0.67 | 0.24 |
| RW | 0.84 | -0.04 | 0.20 |
| SW | -0.37 | 0.04 | 0.90 |
| 特征值Eigenvalue | 2.09 | 1.81 | 0.93 |
| 贡献率/%Contribution rate/% | 34.85 | 30.12 | 15.52 |
| 累计贡献率/% | 34.85 | 64.97 | 80.49 |
| Cumulative contribution rate/% |
| 性状 Trait | 主成分1 PC1 | 主成分2 PC2 | 主成分3 PC3 |
|---|---|---|---|
| PL | 0.76 | -0.19 | 0.52 |
| RL | 0.94 | 0.25 | -0.16 |
| RL/PL | 0.55 | -0.30 | -0.14 |
| GR | 0.65 | 0.53 | -0.51 |
| RW | 0.62 | -0.48 | 0.20 |
| SW | 0.05 | 0.70 | 0.66 |
| 特征值Eigenvalue | 2.56 | 1.20 | 1.05 |
| 贡献率/%Contribution rate/% | 42.68 | 20.01 | 17.51 |
| 累计贡献率/% | 42.68 | 62.69 | 80.20 |
| Cumulative contribution rate/% |
Table 4 Contribution of variables to principal components under cadmium stress across 50 soybean germplasms
| 性状 Trait | 主成分1 PC1 | 主成分2 PC2 | 主成分3 PC3 |
|---|---|---|---|
| PL | 0.76 | -0.19 | 0.52 |
| RL | 0.94 | 0.25 | -0.16 |
| RL/PL | 0.55 | -0.30 | -0.14 |
| GR | 0.65 | 0.53 | -0.51 |
| RW | 0.62 | -0.48 | 0.20 |
| SW | 0.05 | 0.70 | 0.66 |
| 特征值Eigenvalue | 2.56 | 1.20 | 1.05 |
| 贡献率/%Contribution rate/% | 42.68 | 20.01 | 17.51 |
| 累计贡献率/% | 42.68 | 62.69 | 80.20 |
| Cumulative contribution rate/% |
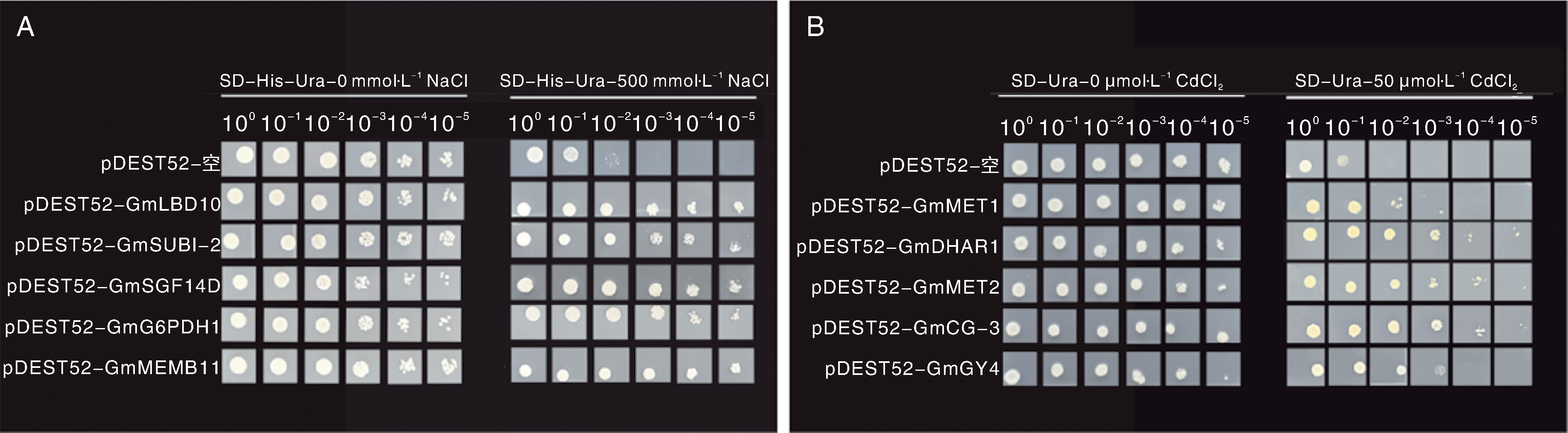
Fig.2 Growth performance of yeast strains transformed with soybean library under different growth conditions A, The survival performance of yeast strain G19 transformed with full-length soybean cDNA library on SD-His-Ura medium supplemented with 500 mmol·L-1 NaCl; B, The survival performance of yeast strain Δycf1 transformed with full-length soybean cDNA library on SD-Ura medium supplemented with 50 μmol·L-1 CdCl2. The photos were taken three days after plating.
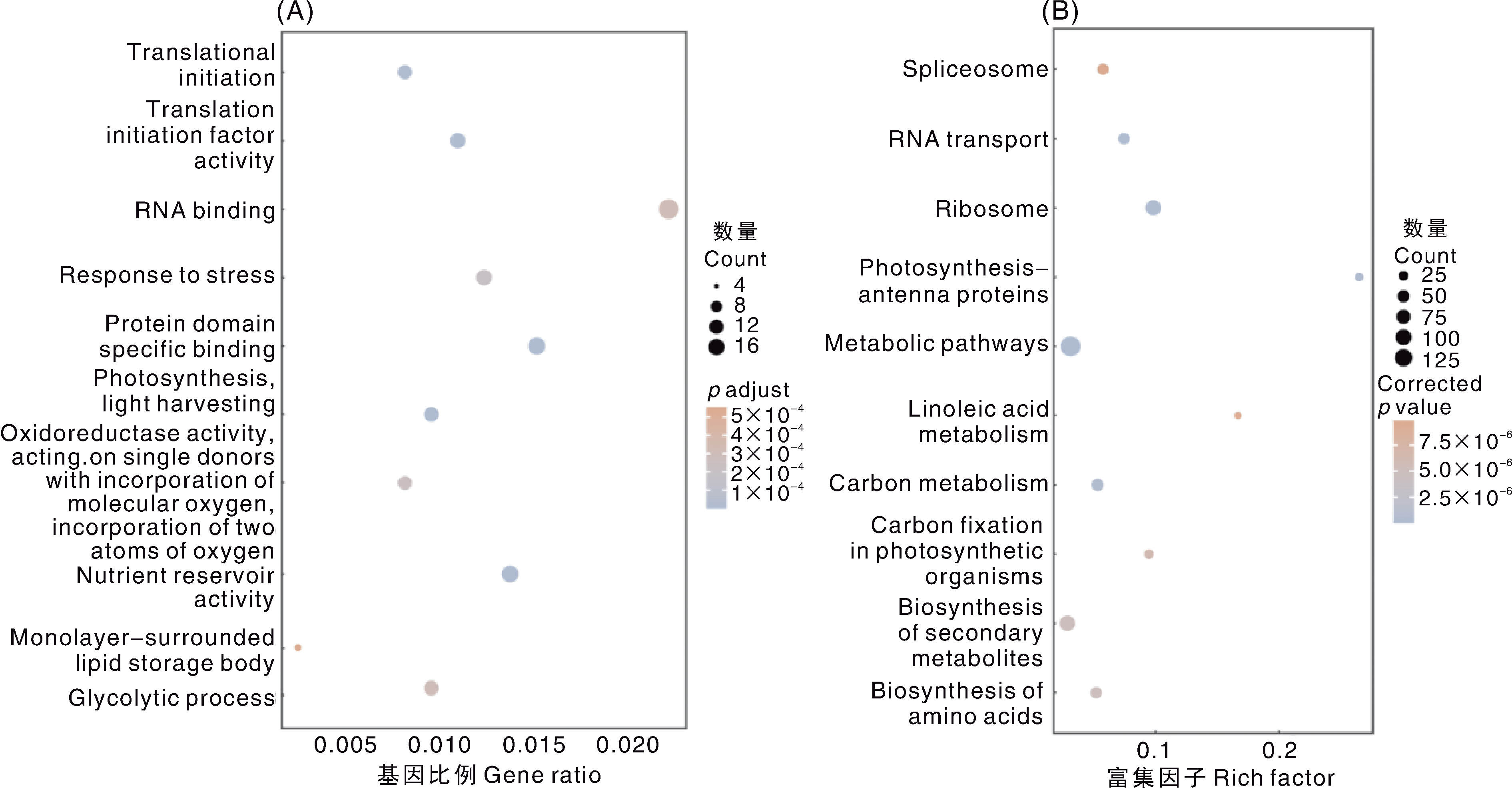
Fig.3 GO and KEGG analysis of candidate genes for salt tolerance in soybean A, GO function clustering analysis; B, KEGG pathway enrichment analysis. The annotation names of Fig. A from top to bottom are as follows: Translational initiation; Translation initiation factor activity; RNA binding; Response to stress; Protein domain specific binding; Photosynthesis, light harvesting; Oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen; Nutrient reservoir activity; Monolayer-surrounded lipid storage body; Glycolytic process. The annotation names of Fig. B from top to bottom are as follows: Spliceosome; RNA transport; Ribosome; Photosynthesis-antenna proteins; Metabolic pathways; Linoleic acid metabolism; Carbon metabolism; Carbon fixation in photosynthetic organisms; Biosynthesis of secondary metabolites; Biosynthesis of amino acids.
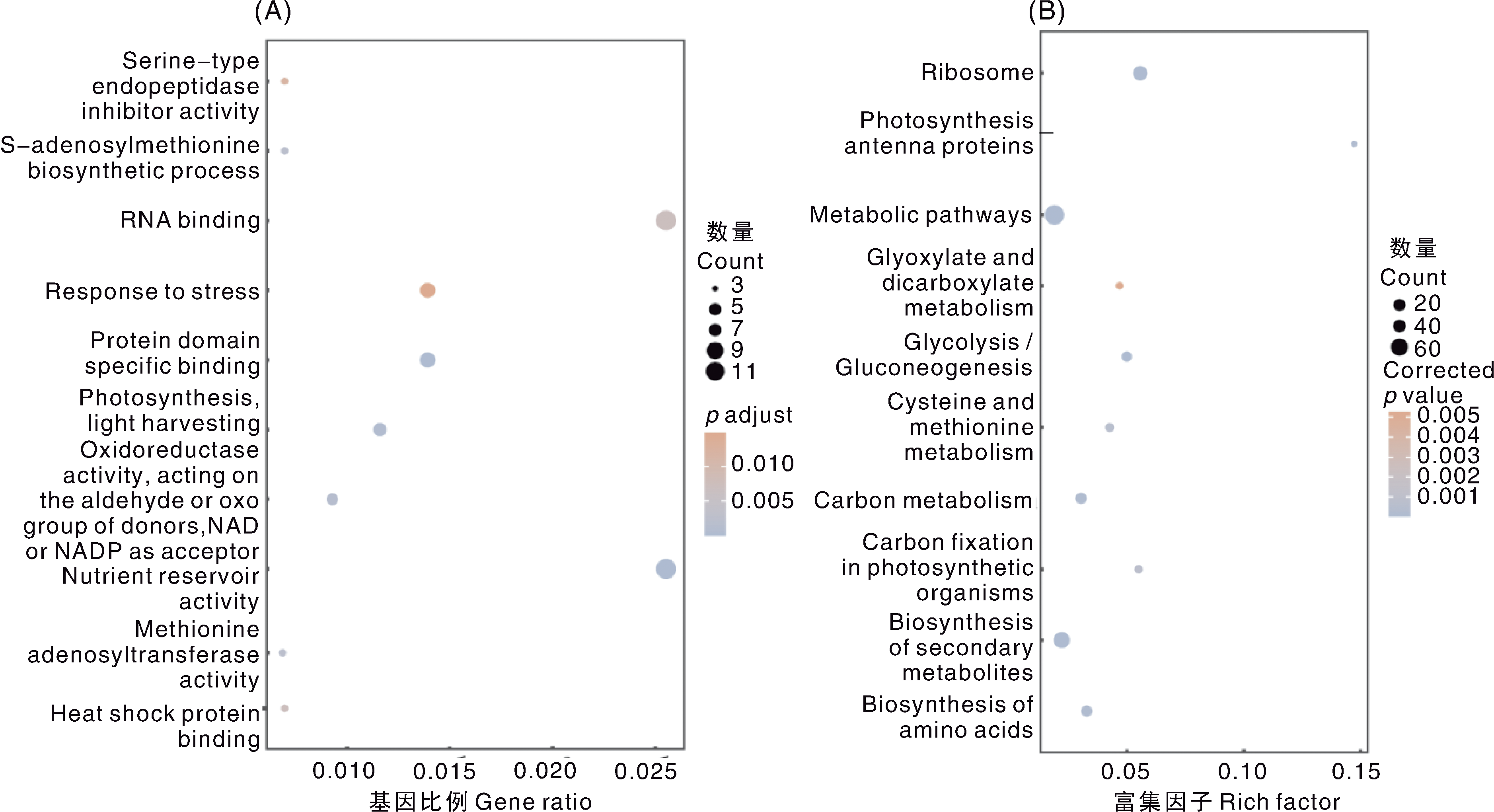
Fig.4 GO and KEGG enrichment analysis of candidate genes for cadmium tolerance in soybean A, GO function clustering analysis; B, KEGG pathway enrichment analysis. The annotation names of Fig. A from top to bottom are as follows: Serine-type endopeptidase inhibitor activity; S-adenosylmethionine biosynthetic process; RNA binding; Response to stress; Protein domain specific binding; Photosynthesis, light harvesting; Oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor; Nutrient reservoir activity; Methionine adenosyltransferase activity; Heat shock protein binding. The annotation names of Fig. B from top to bottom are as follows: Ribosome; Photosynthesis-antenna proteins; Metabolic pathways; Glyoxylate and dicarboxylate metabolism; Glycolysis/Gluconeogenesis; Cysteine and methionine metabolism; Carbon metabolism; Carbon fixation in photosynthetic organisms; Biosynthesis of secondary metabolites; Biosynthesis of amino acids.
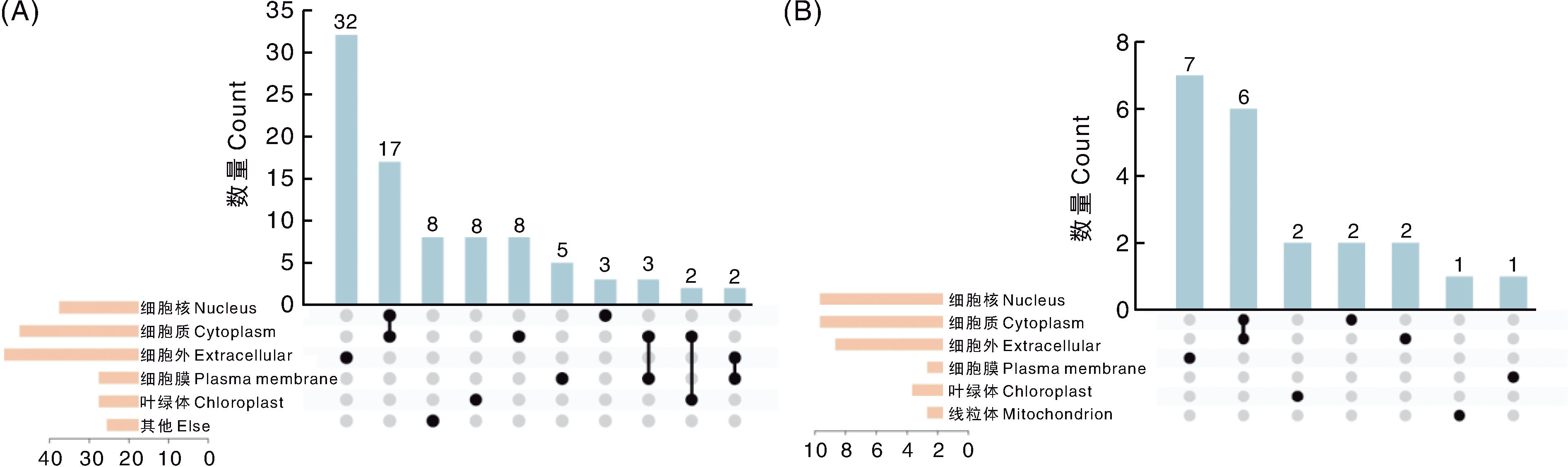
Fig.5 Subcellular localization of proteins encoded by soybean salt and cadmium tolerance candidate genes A, Subcellular localization of proteins encoded by known genes; B, Subcellular localization of proteins encoded by unknown genes.
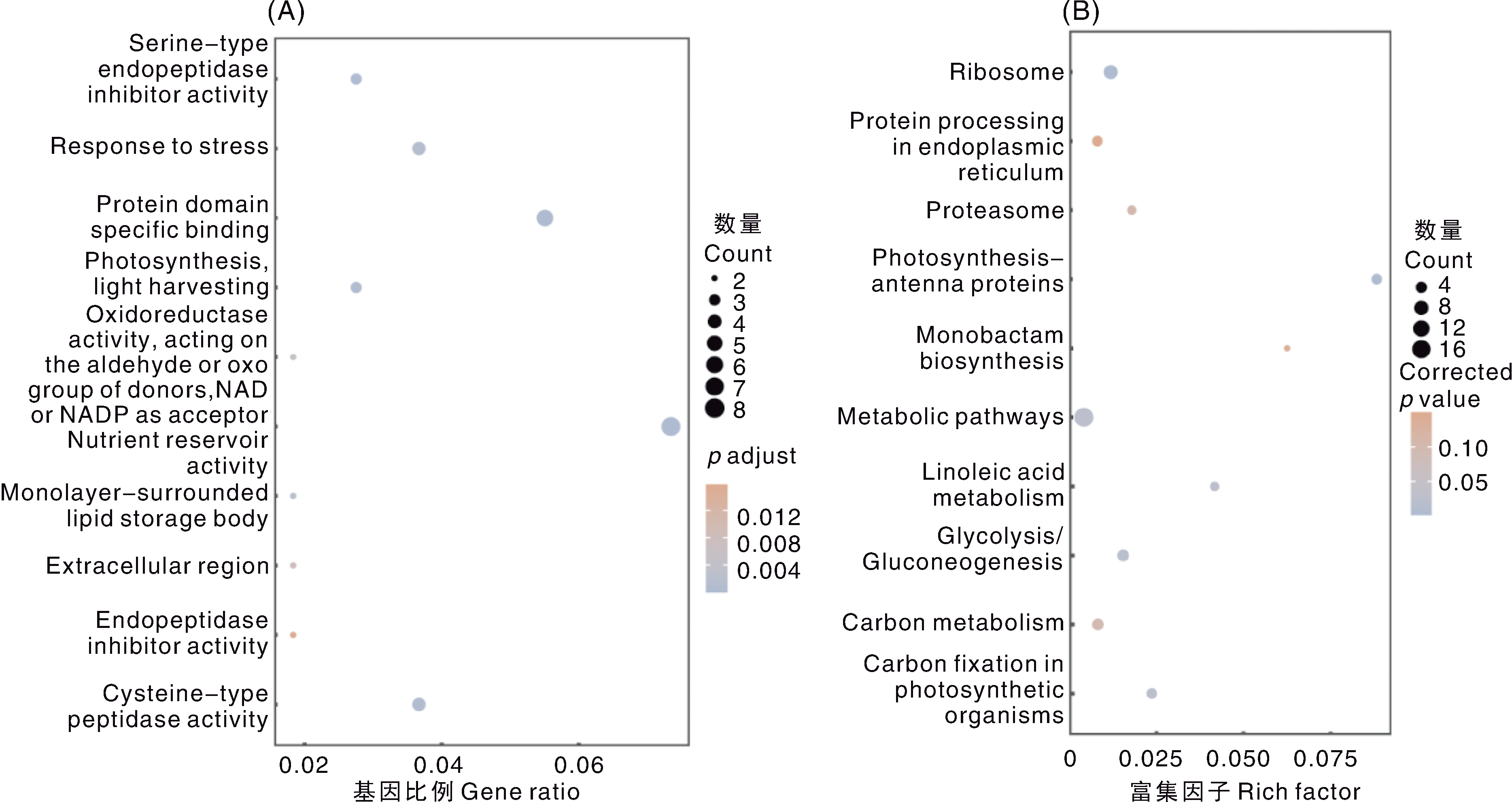
Fig.6 GO and KEGG analysis of candidate genes for salt-cadmium co-tolerance in soybean A, GO function clustering analysis; B, KEGG pathway enrichment analysis. The annotation names of Fig. A from top to bottom are as follows: Serine-type endopeptidase inhibitor activity; Response to stress; Protein domain specific binding; Photosynthesis, light harvesting; Oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor; Nutrient reservoir activity; Monolayer-surrounded lipid storage body; Extracellular region; Endopeptidase inhibitor activity; Cysteine-type peptidase activity. The annotation names of Fig. B from top to bottom are as follows: Ribosome; Protein processing in endoplasmic reticulum; Proteasome; Photosynthesis-antenna proteins; Monobactam biosynthesis; Metabolic pathways; Linoleic acid metabolism; Glycolysis/Gluconeogenesis; Carbon metabolism; Carbon fixation in photosynthetic organisms. Gene ratio represents the ratio of differentially expressed genes annotated to a specific term to the total number of differentially expressed genes; Rich factor is the ratio of enriched differentially expressed genes in a pathway to all genes in that pathway. The same as below.
| 基因ID Gene ID | 基因名 Gene name | cDNA大小/bp cDNA size/bp | 预测功能 Function prediction |
|---|---|---|---|
| Glyma.09G185500 | GmDehydrin | 681 | Dehydrin-like protein |
| Glyma.03G163533 | GmSSP | 1 488 | Rmlc-like cupins superfamily protein |
| Glyma.02G208700 | GmGF14 | 789 | 14-3-3-like protein |
| Glyma.10G246300 | GmPAP85 | 1 866 | Cupin family protein |
| Glyma.15G072400 | GmHUP54 | 756 | Aluminium induced protein with YGL and LRDR motifs |
| Glyma.07G132000 | GmMET2 | 240 | Metallothionein 2A |
Table 5 Key salt and cadmium tolerant gene information of soybean
| 基因ID Gene ID | 基因名 Gene name | cDNA大小/bp cDNA size/bp | 预测功能 Function prediction |
|---|---|---|---|
| Glyma.09G185500 | GmDehydrin | 681 | Dehydrin-like protein |
| Glyma.03G163533 | GmSSP | 1 488 | Rmlc-like cupins superfamily protein |
| Glyma.02G208700 | GmGF14 | 789 | 14-3-3-like protein |
| Glyma.10G246300 | GmPAP85 | 1 866 | Cupin family protein |
| Glyma.15G072400 | GmHUP54 | 756 | Aluminium induced protein with YGL and LRDR motifs |
| Glyma.07G132000 | GmMET2 | 240 | Metallothionein 2A |
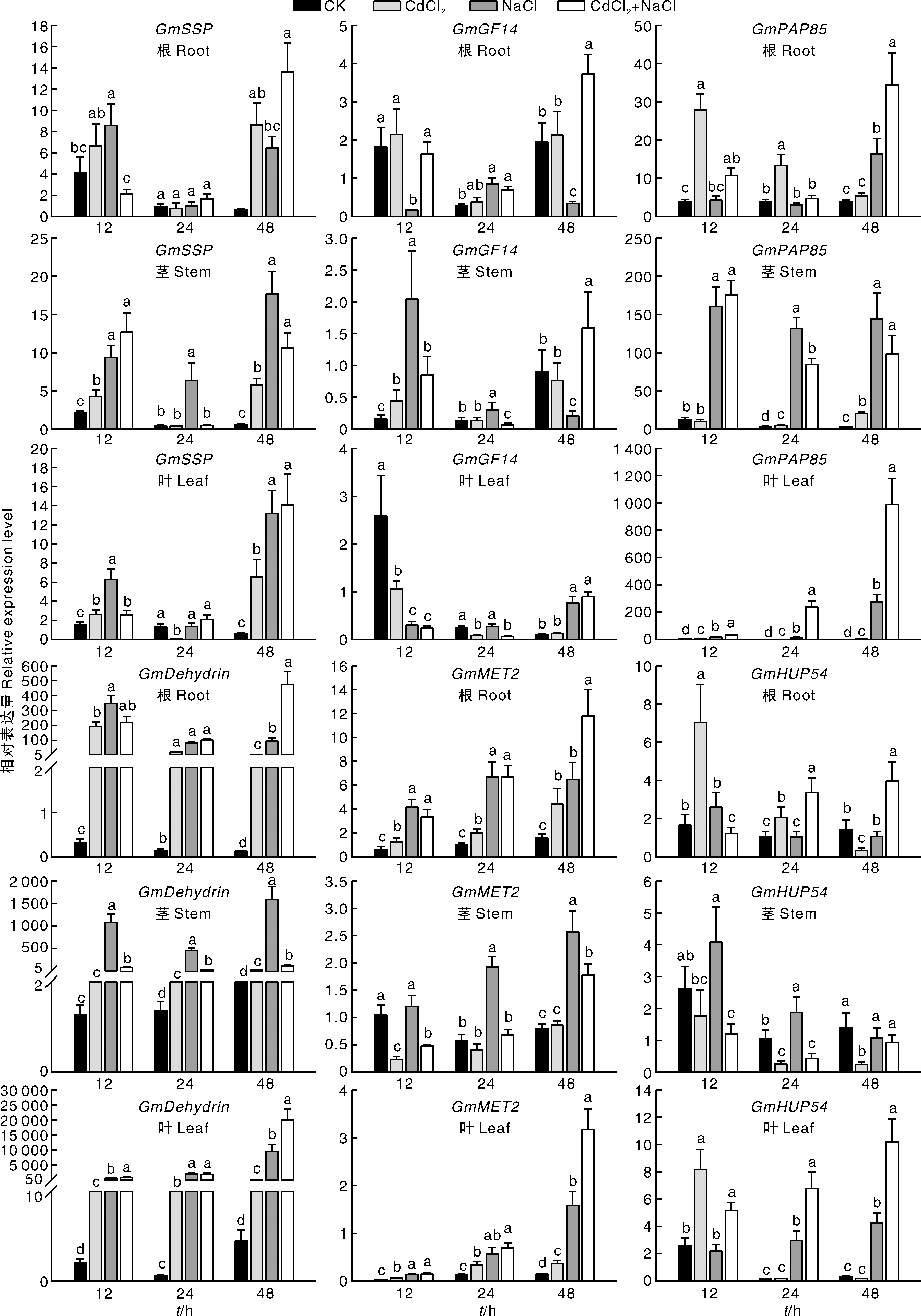
Fig.7 Relative expression level of candidate genes in soybean roots, stems, and leaves under salt, Cd and salt-Cd stress Bars marked with the same lowercase letters in the figure indicate significant differences (p<0.05) among treatments at the same time.
| [1] | HAMZA M, BASIT A W, SHEHZADI I, et al. Global impact of soybean production: a review[J]. Asian Journal of Biochemistry, Genetics and Molecular Biology, 2024, 16(2): 12-20. |
| [2] | 李奕聪, 杨钰莹, 李佳璇, 等. 2024年大豆产业发展趋势与政策建议[J]. 大豆科技, 2024(1): 1-5. |
| LI Y C, YANG Y Y, LI J X, et al. Development trends and policy suggestions of soybean industry in 2024[J]. Soybean Science & Technology, 2024(1): 1-5. | |
| [3] | 闫琰, 王秀东, 王济民, 等. “双循环” 背景下国家粮食安全战略研究[J]. 中国工程科学, 2023, 25(4): 14-25. |
| YAN Y, WANG X D, WANG J M, et al. National food security strategy against the backdrop of domestic and international dual circulation[J]. Strategic Study of CAE, 2023, 25(4): 14-25. | |
| [4] | SHANG C L, WANG L, TIAN C Y, et al. Heavy metal tolerance and potential for remediation of heavy metal-contaminated saline soils for the euhalophyte Suaeda salsa[J]. Plant Signaling & Behavior, 2020, 15(11): 1805902. |
| [5] | 唐希望, 周阳, 王龙雪, 等. 盐镉耦合胁迫对玉米种子萌发和幼苗生长的影响[J]. 东北农业科学, 2022, 47(5): 25-29. |
| TANG X W, ZHOU Y, WANG L X, et al. Effects of salt-cadmium coupling stress on seed germination and seedling growth of maize[J]. Journal of Northeast Agricultural Sciences, 2022, 47(5): 25-29. | |
| [6] | ZHANG W, LIAO X L, CUI Y M, et al. A cation diffusion facilitator, GmCDF1, negatively regulates salt tolerance in soybean[J]. PLoS Genetics, 2019, 15(1): e1007798. |
| [7] | SINGLETON P W, BEN BOHLOOL B. Effect of salinity on nodule formation by soybean[J]. Plant Physiology, 1984, 74(1): 72-76. |
| [8] | EL-SABAGH A, SOROUR S G R, UEDA A, et al. Evaluation of salinity stress effects on seed yield and quality of three soybean cultivars[J]. Azarian Journal of Agriculture, 2015, 2(5): 138-141. |
| [9] | DO T D, VUONG T D, DUNN D, et al. Mapping and confirmation of loci for salt tolerance in a novel soybean germplasm, Fiskeby Ⅲ[J]. Theoretical and Applied Genetics, 2018, 131(3): 513-524. |
| [10] | 黄运湘, 廖柏寒, 肖浪涛, 等. 镉处理对大豆幼苗生长及激素含量的影响[J]. 环境科学, 2006, 27(7): 1398-1401. |
| HUANG Y X, LIAO B H, XIAO L T, et al. Effects of Cd2+ on seedling growth and phytohormone contents of Glycine max[J]. Environmental Science, 2006, 27(7): 1398-1401. | |
| [11] | 陈朝明, 龚惠群, 王凯荣. Cd对桑叶品质、生理生化特性的影响及其机理研究[J]. 应用生态学报, 1996, 7(4): 417-423. |
| CHEN C M, GONG H Q, WANG K R. Effect of Cd on quality, physiological and biochemical characteristics of mulberry leaves and its mechanism[J]. Chinese Journal of Applied Ecology, 1996, 7(4): 417-423. | |
| [12] | LI J, GUO Y, YANG Y Q. The molecular mechanism of plasma membrane H+-ATPases in plant responses to abiotic stress[J]. Journal of Genetics and Genomics, 2022, 49(8): 715-725. |
| [13] | VAN ZELM E, ZHANG Y X, TESTERINK C. Salt tolerance mechanisms of plants[J]. Annual Review of Plant Biology, 2020, 71: 403-433. |
| [14] | FENG C, GAO H T, ZHOU Y G, et al. Unfolding molecular switches for salt stress resilience in soybean: recent advances and prospects for salt-tolerant smart plant production[J]. Frontiers in Plant Science, 2023, 14: 1162014. |
| [15] | AHMAD I, MIAN A, MAATHUIS F J M. Overexpression of the rice AKT1 potassium channel affects potassium nutrition and rice drought tolerance[J]. Journal of Experimental Botany, 2016, 67(9): 2689-2698. |
| [16] | DEINLEIN U, STEPHAN A B, HORIE T, et al. Plant salt-tolerance mechanisms[J]. Trends in Plant Science, 2014, 19(6): 371-379. |
| [17] | CHUNG Y S, KIM K S, HAMAYUN M, et al. Silicon confers soybean resistance to salinity stress through regulation of reactive oxygen and reactive nitrogen species[J]. Frontiers in Plant Science, 2020, 10: 1725. |
| [18] | 孙敏. 水稻植株中镉区室化关键螯合物的鉴定与分析[D]. 北京: 中国农业科学院, 2010. |
| SUN M. Characterization and analysis of chelates involved in compartmentation of cadmium in rice plants[D]. Beijing: Chinese Academy of Agricultural Sciences, 2010. | |
| [19] | 张旭红, 高艳玲, 林爱军, 等. 植物根系细胞壁在提高植物抵抗金属离子毒性中的作用[J]. 生态毒理学报, 2008, 3(1): 9-14. |
| ZHANG X H, GAO Y L, LIN A J, et al. A review on the effects of cell wall on the resistance of plants to metal stress[J]. Asian Journal of Ecotoxicology, 2008, 3(1): 9-14. | |
| [20] | HIRATA K, TSUJI N, MIYAMOTO K. Biosynthetic regulation of phytochelatins, heavy metal-binding peptides[J]. Journal of Bioscience and Bioengineering, 2005, 100(6): 593-599. |
| [21] | LUO P, WU J J, LI T T, et al. An overview of the mechanisms through which plants regulate ROS homeostasis under cadmium stress[J]. Antioxidants, 2024, 13(10): 1174. |
| [22] | FALLER P, KIENZLER K, KRIEGER-LISZKAY A. Mechanism of Cd2+ toxicity: Cd2+ inhibits photoactivation of Photosystem Ⅱ by competitive binding to the essential Ca2+ site[J]. Biochimica et Biophysica Acta(BBA)-Bioenergetics 2005, 1706(1/2): 158-164. |
| [23] | EL-SHINTINAWY F. Glutathione counteracts the inhibitory effect induced by cadmium on photosynthetic process in soybean[J]. Photosynthetica, 1999, 36(1): 171-179. |
| [24] | NING H X, ZHANG C H, YAO Y, et al. Overexpression of a soybean O-acetylserine (thiol) lyase-encoding gene GmOASTL4 in tobacco increases cysteine levels and enhances tolerance to cadmium stress[J]. Biotechnology Letters, 2010, 32(4): 557-564. |
| [25] | AZEVEDO R A, GRATÃO P L, MONTEIRO C C, et al. What is new in the research on cadmium-induced stress in plants?[J]. Food and Energy Security, 2012, 1(2): 133-140. |
| [26] | PESSARAKLI M. Handbook of plant and crop physiology[M]. 4th ed. Boca Raton: CRC Press, 2021. |
| [27] | UDDIN K M, JURAIMI A S, ISMAIL M R, et al. Relative salinity tolerance of warm season turfgrass species[J]. Journal of Environmental Biology, 2011, 32(3): 309-312. |
| [28] | CHEN Y, LI L L, ZONG J Q, et al. Heterologous expression of the halophyte Zoysia matrella H+-pyrophosphatase gene improved salt tolerance in Arabidopsis thaliana[J]. Plant Physiology and Biochemistry, 2015, 91: 49-55. |
| [29] | DALCORSO G, FARINATI S, FURINI A. Regulatory networks of cadmium stress in plants[J]. Plant Signaling & Behavior, 2010, 5(6): 663-667. |
| [30] | 韩毅强, 高亚梅, 杜艳丽, 等. 大豆耐盐碱种质资源鉴定[J]. 中国油料作物学报, 2021, 43(6): 1016-1024. |
| HAN Y Q, GAO Y M, DU Y L, et al. Identification of saline-alkali tolerant germplasm resources of soybean during the whole growth stage[J]. Chinese Journal of Oil Crop Sciences, 2021, 43(6): 1016-1024. | |
| [31] | 林峰, 赵慧艳, 史飞飞, 等. 大豆种质资源苗期耐盐鉴定及遗传多样性分析[J]. 植物遗传资源学报, 2024, 25(6): 945-956. |
| LIN F, ZHAO H Y, SHI F F, et al. Identification of salt-tolerant germplasm resources in soybean seedlings and genetic diversity analysis[J]. Journal of Plant Genetic Resources, 2024, 25(6): 945-956. | |
| [32] | ARAO T, ISHIKAWA S. Genotypic differences in cadmium concentration and distribution of soybean and rice[J]. Japan Agricultural Research Quarterly, 2006, 40(1): 21-30. |
| [33] | 周秀文, 张晓蕊, 孙贺祥, 等. 大豆种质萌发期和苗期耐盐性评价[J]. 沈阳农业大学学报, 2022, 53(3): 257-264. |
| ZHOU X W, ZHANG X R, SUN H X, et al. Evaluation of salt tolerance of soybean germplasms at germination and seedling stages[J]. Journal of Shenyang Agricultural University, 2022, 53(3): 257-264. | |
| [34] | 严勇亮, 张金波, 路子峰, 等. 大豆种质资源耐盐性鉴定与评价[J]. 新疆农业科学, 2021, 58(1): 65-71. |
| YAN Y L, ZHANG J B, LU Z F, et al. Salt tolerance evaluation of soybean germplasm[J]. Xinjiang Agricultural Sciences, 2021, 58(1): 65-71. | |
| [35] | SHAMSI I H, ZHANG G P, HU H L, et al. Assessment of the hazardous effects of Cd on physiological and biochemical characteristics of soybean genotypes[J]. International Journal of Agriculture and Biology, 2014, 16(1): 41-48. |
| [36] | ZHI Y, SUN T, ZHOU Q X, et al. Screening of safe soybean cultivars for cadmium contaminated fields[J]. Scientific Reports, 2020, 10: 12965. |
| [37] | 石广成. 大豆耐盐种质筛选及GmSCAMPs家族的耐盐功能分析[D]. 太谷: 山西农业大学, 2022. |
| SHI G C. Screening of soybean salt-tolerant germplasm and study on salt-tolerant function of GmSCAMPs family[D]. Taigu: Shanxi Agricultural University, 2022. | |
| [38] | 袁宇婷, 张晓燕, 吴谷丰, 等. 基于主成分和隶属函数分析的大豆种质资源耐盐性综合评价[J]. 大豆科学, 2025, 44(1): 22-32. |
| YUAN Y T, ZHANG X Y, WU G F, et al. Comprehensive evaluation of salt tolerance of soybean germplasm resources based on principal component and membership function analysis[J]. Soybean Science, 2025, 44(1): 22-32. | |
| [39] | 张新草, 薛项潇, 姜深, 等. 大豆种质发芽期耐盐碱性鉴定及指标筛选[J]. 西北农业学报, 2020, 29(3): 374-381. |
| ZHANG X C, XUE X X, JIANG S, et al. Identification of mixed saline-alkali tolerance and screening of indicators in soybean at germination stage[J]. Acta Agriculturae Boreali-occidentalis Sinica, 2020, 29(3): 374-381. | |
| [40] | 刘佳丽. 重金属镉对大豆种子萌发与幼苗生长的影响[J]. 农业技术与装备, 2023(12): 13-15. |
| LIU J L. Effects of heavy metal cadmium on seed germination and seedling growth of soybean[J]. Agricultural Technology & Equipment, 2023(12): 13-15. | |
| [41] | 郭远, 王文成, 徐颖莹, 等. 植物耐盐评价方法综述[J]. 江苏农业科学, 2017, 45(23): 18-23. |
| GUO Y, WANG W C, XU Y Y, et al. Review on evaluation methods of plant salt tolerance[J]. Jiangsu Agricultural Sciences, 2017, 45(23): 18-23. | |
| [42] | TAO J Y, LU L L, TAO J Y, et al. Advances in genes-encoding transporters for cadmium uptake, translocation, and accumulation in plants[J]. Toxics, 2022, 10(8): 411. |
| [43] | SHIRAKU M L, MAGWANGA R O, ZHANG Y Y, et al. Late embryogenesis abundant gene LEA3(Gh_A08G0694) enhances drought and salt stress tolerance in cotton[J]. International Journal of Biological Macromolecules, 2022, 207: 700-714. |
| [44] | LI X, FENG H, LIU S, et al. Dehydrin CaDHN2 enhances drought tolerance by affecting ascorbic acid synthesis under drought in peppers[J]. Plants, 2023, 12(22): 3895. |
| [45] | PAL P, MASAND M, SHARMA S, et al. Genome-wide transcriptional profiling and physiological investigation elucidating the molecular mechanism of multiple abiotic stress response in Stevia rebaudiana Bertoni[J]. Scientific Reports, 2023, 13: 19853. |
| [46] | SHIRAKU M L, MAGWANGA R O, CAI X Y, et al. Knockdown of 60S ribosomal protein L14-2 reveals their potential regulatory roles to enhance drought and salt tolerance in cotton[J]. Journal of Cotton Research, 2021, 4(1): 27. |
| [47] | GONG J W, LIU Q, CAI L L, et al. Multimechanism collaborative superior antioxidant CDzymes to alleviate salt stress-induced oxidative damage in plant growth[J]. ACS Sustainable Chemistry & Engineering, 2023, 11(10): 4237-4247. |
| [48] | CHUNG E S, KIM K Y, SO H A, et al. Enhanced tolerance against osmotic stresses of Escherichia coli cells expressing soybean KS-type dehydrin[J]. Genes & Genomics, 2008, 30(4): 319-327. |
| [49] | ŠVECOVÁ M, BOSZORÁDOVÁ E, MATUŠÍKOVÁ I, et al. Arabidopsis AtLTI30 and AtHIRD11 dehydrin genes and their contribution to cadmium tolerance in transgenic tobacco plants[J]. Acta Physiologiae Plantarum, 2022, 45(2): 21. |
| [50] | AHAMMED G J, LI C X, LI X, et al. Overexpression of tomato RING E3 ubiquitin ligase gene SlRING1 confers cadmium tolerance by attenuating cadmium accumulation and oxidative stress[J]. Physiologia Plantarum, 2021, 173(1): 449-459. |
| [51] | LIU J, QI W C, LU H Y, et al. Characterization of interactions between the soybean salt-stress responsive membrane-intrinsic proteins GmPIP1 and GmPIP2[J]. Agronomy, 2021, 11(7): 1312. |
| [52] | ZHOU L, WANG C, LIU R F, et al. Constitutive overexpression of soybean plasma membrane intrinsic protein GmPIP1;6 confers salt tolerance[J]. BMC Plant Biology, 2014, 14: 181. |
| [53] | HU W, YUAN Q Q, WANG Y, et al. Overexpression of a wheat aquaporin gene, TaAQP8, enhances salt stress tolerance in transgenic tobacco[J]. Plant & Cell Physiology, 2012, 53(12): 2127-2141. |
| [54] | PAGANI M A, TOMAS M, CARRILLO J, et al. The response of the different soybean metallothionein isoforms to cadmium intoxication[J]. Journal of Inorganic Biochemistry, 2012, 117: 306-315. |
| [55] | MANOSALVA P M, BRUCE M, LEACH J E. Rice 14-3-3 protein (GF14e) negatively affects cell death and disease resistance[J]. Plant Journal, 2011, 68(5): 777-787. |
| [56] | QIU W M, SONG X X, HAN X J, et al. Overexpression of Sedum alfredii cinnamyl alcohol dehydrogenase increases the tolerance and accumulation of cadmium in Arabidopsis[J]. Environmental and Experimental Botany, 2018, 155: 566-577. |
| [57] | ZHANG J, YANG N, LI Y Y, et al. Overexpression of PeMIPS1 confers tolerance to salt and copper stresses by scavenging reactive oxygen species in transgenic poplar[J]. Tree Physiology, 2018, 38(10): 1566-1577. |
| [1] | HU Yingjie, DU Chenqi, WANG Liufan, SHOU Jianxin, WANG Chao, XU Mei, YAN Xu. Research progress of vesicle trafficking in plant response to salt stress [J]. Acta Agriculturae Zhejiangensis, 2025, 37(9): 2003-2011. |
| [2] | GUAN Xiusheng, LIU Tieshan, WANG Juan, ZHANG Maolin, LIU Chunxiao, DONG Rui, GUAN Haiying, LIU Qiang, XU Yang, HE Chunmei. Bioinformatics analysis and cloning of NF-YA family genes in maize(Zea mays) [J]. Acta Agriculturae Zhejiangensis, 2025, 37(8): 1605-1614. |
| [3] | HE Guoxin, LI Sujuan, WANG Jian, TAO Xiaoyuan, YE Zihong, CHEN Guang, XU Shengchun. Screening and identification of soybean germplasm for low nitrogen tolerance during seedling stage [J]. Acta Agriculturae Zhejiangensis, 2025, 37(5): 965-976. |
| [4] | XU Zhuwei, LEI Jun, SHAO Xiaowei, CHEN Runxing, JIANG Huan, WANG Shougen, YU Wenhui. Evaluation of Quzhou fresh soybean oligosaccharide germplasm resources based on analytic hierarchy process and fuzzy comprehensive evaluation method [J]. Acta Agriculturae Zhejiangensis, 2025, 37(4): 754-766. |
| [5] | TANG Aoran, JIN Xiu, WANG Tan, RAO Yuan, LI Jiajia, ZHANG Wu. Physiological plant height measurement method based on the reconstruction of the main stem skeleton for curved soybean plants [J]. Acta Agriculturae Zhejiangensis, 2025, 37(2): 466-479. |
| [6] | LIAO Xiaolong, WANG Xingsheng, CHEN Yong, LI Bin, HONG Sidan, MEI Lina, GUO Ying. Identification of the HKT gene family members in Populus species and analysis of their expression patterns under salt stress [J]. Acta Agriculturae Zhejiangensis, 2025, 37(10): 2104-2115. |
| [7] | OU Jinwen, ZHANG Guwen, FENG Zhijuan, WANG Bin, BU Yuanpeng, XU Yu, RU Lei, LIU Na, GONG Yaming. Identification of soybean trehalose-6-phosphate phosphatase gene GmTPP and its expression analysis in growth and abiotic stress response [J]. Acta Agriculturae Zhejiangensis, 2024, 36(9): 2031-2041. |
| [8] | PENG Jiacheng, WU Yue, XU Jiehao, XIA Meiwen, QI Tianpeng, XU Haisheng. Cloning of paxillin gene from Macrobrachium nipponense and effect of cadmium stress on its expression [J]. Acta Agriculturae Zhejiangensis, 2024, 36(2): 247-253. |
| [9] | GAO Jing, LU Linghong, GU Xianbin, FAN Fei, SONG Genhua, ZHANG Huiqin. Cloning of AcWRKY94 gene from kiwifruit and its functional analysis under salt stress [J]. Acta Agriculturae Zhejiangensis, 2024, 36(11): 2501-2509. |
| [10] | TANG Yuehui, CHEN Shuying, HE Wenqiong, WANG Hanjin, BAO Xinxin, JIA Sainan, WANG Yaoyao, CHEN Yuyang, YANG Tongwen. Cloning and functional analysis of JcERF22 gene from Jatropha curcas [J]. Acta Agriculturae Zhejiangensis, 2024, 36(10): 2219-2228. |
| [11] | SUN Xiujuan, XU Weihui, WANG Zhigang. Isolation and identification of endophytic bacteria from soybean nodule and their effects on soybean plant [J]. Acta Agriculturae Zhejiangensis, 2023, 35(7): 1532-1541. |
| [12] | BU Yuanpeng, LIU Na, ZHANG Guwen, FENG Zhijuan, WANG Bin, GONG Yaming, XU Linying. Diversity evaluation of agronomic traits and construction of core collection and taste quality evaluation system in vegetable soybean germplasm resources [J]. Acta Agriculturae Zhejiangensis, 2023, 35(6): 1307-1314. |
| [13] | YANG Songhua, SHI Guiyang, WANG Jingqin, CHEN Zhu. Effects of soybean root exudates on insoluble phosphorus in soil under low phosphorus stress [J]. Acta Agriculturae Zhejiangensis, 2023, 35(6): 1396-1406. |
| [14] | TAN Shuxia, ZHAO Taodi, YANG Hao, NING Kejun, LIU Li, HE Qingyuan, HUANG Shoucheng, SHU Yingjie. Effects of shading on agronomic characters, yield and nitrogen metabolism of 10 vegetable soybean varieties [J]. Acta Agriculturae Zhejiangensis, 2023, 35(4): 729-735. |
| [15] | ZHANG Meng, SHE Bao, YANG Yuying, HUANG Linsheng, ZHU Mengqi. Study on extraction method of soybean planting areas based on unmanned aerial vehicle RGB image [J]. Acta Agriculturae Zhejiangensis, 2023, 35(4): 952-961. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||

